Difference between revisions of "Welcome to the In-Silico Model of Cutaneous Lipids Wiki"
(→Description of the Model) |
|||
| Line 1: | Line 1: | ||
== Description of the Model == | == Description of the Model == | ||
| − | Computational models have been demonstrated as useful tools to gain a better understanding of networks, aid rational decision-making and support research across the biosciences. Due to the complexity of the AA cascade, predicting the outcome of the network is difficult due to the collaborative action of numerous species (i.e. metabolites, proteins, signalling cascades). In addition, the AA cascade is ubiquitous and is implicated in numerous inflammatory diseases such as cancer, atherosclerosis and asthma; as a result, the production of eicosanoids is the targets of several pharmacological interventions. | + | Computational models have been demonstrated as useful tools to gain a better understanding of networks, aid rational decision-making and support research across the biosciences. Due to the complexity of the AA cascade, predicting the outcome of the network is difficult due to the collaborative action of numerous species (i.e. metabolites, proteins, signalling cascades). In addition, the AA cascade is ubiquitous and is implicated in numerous inflammatory diseases such as cancer, atherosclerosis and asthma (Bishop-Bailey et al., 2002, von Schacky, 2015, Claar et al., 2015, Albanesi et al., 2010); as a result, the production of eicosanoids is the targets of several pharmacological interventions. |
Therefore, there is a need to utilise a systems-level approach to further investigate the bioactive AA cascade. This can be achieved by building upon previous models of the AA cascade and creating a new model which addresses the gaps in the literature such as an extensive network topology, the use of experimental parameters, documentation of parameter sources, integration of data from multiple platforms and application beyond a single cell type. | Therefore, there is a need to utilise a systems-level approach to further investigate the bioactive AA cascade. This can be achieved by building upon previous models of the AA cascade and creating a new model which addresses the gaps in the literature such as an extensive network topology, the use of experimental parameters, documentation of parameter sources, integration of data from multiple platforms and application beyond a single cell type. | ||
Revision as of 14:54, 20 August 2019
Contents
Description of the Model
Computational models have been demonstrated as useful tools to gain a better understanding of networks, aid rational decision-making and support research across the biosciences. Due to the complexity of the AA cascade, predicting the outcome of the network is difficult due to the collaborative action of numerous species (i.e. metabolites, proteins, signalling cascades). In addition, the AA cascade is ubiquitous and is implicated in numerous inflammatory diseases such as cancer, atherosclerosis and asthma (Bishop-Bailey et al., 2002, von Schacky, 2015, Claar et al., 2015, Albanesi et al., 2010); as a result, the production of eicosanoids is the targets of several pharmacological interventions.
Therefore, there is a need to utilise a systems-level approach to further investigate the bioactive AA cascade. This can be achieved by building upon previous models of the AA cascade and creating a new model which addresses the gaps in the literature such as an extensive network topology, the use of experimental parameters, documentation of parameter sources, integration of data from multiple platforms and application beyond a single cell type.
Furthermore, there is a need to incorporate novel approaches to the parameterisation and simulation of metabolic models, in order to quantify the uncertainty associated with literature data and indicate the robustness of each prediction.
The main aim of this project is to construct, analyse and validate a predictive and adaptable metabolic model of the AA cascade. To use this model to investigate how the concentration of metabolites change over time following internal/external perturbations, we need to utilise a kinetic deterministic modelling approach. This work will build upon the previous models of the AA cascade and support researchers across the biosciences to gain a better understanding of pathways by incorporating means of adapting the model to represent different cell types. We will demonstrate the application of this work in immortalised human cell lines, HaCaT keratinocytes and 46BR.1N fibroblasts as they are shown to produce different eicosanoids, availabilities of AA, protein profiles and response to inflammatory stimuli.
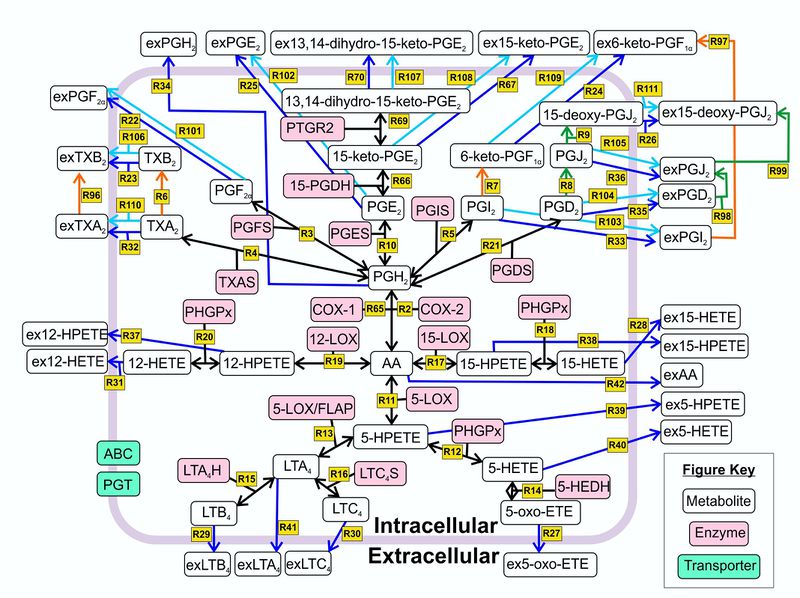
Reactions
Literature information surrounding every reaction within the model are described on the following pages: