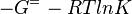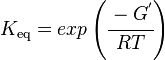Difference between revisions of "Kinetic Model of Monoterpenoid Biosynthesis Wiki"
Aliah.hawari (talk | contribs) (→Description of the model) |
Aliah.hawari (talk | contribs) (→Description of the model) |
||
| Line 10: | Line 10: | ||
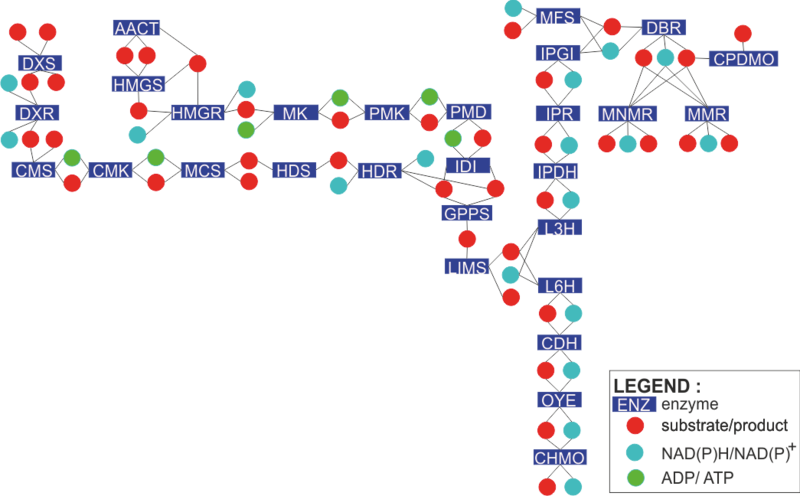
File:Network_v2.png|center| 800px| Schematic representation of the monoterpenoid biosynthesis network being modelled | File:Network_v2.png|center| 800px| Schematic representation of the monoterpenoid biosynthesis network being modelled | ||
| − | rect 583 336 639 358 [[Limonene | + | rect 579 265 643 287 [[ Geranyl diphosphate synthase (GPPS)]] |
| − | rect | + | rect 583 336 639 358 [[Limonene synthase]] |
| − | rect | + | rect 839 25 897 44 [[Pulegone Reductase (PGR)]] |
| − | rect | + | rect 782 137 858 158 [[ Menthone:Neomenthol reductase (MNMR)]] |
| − | rect | + | rect 897 138 954 157 [[ Menthone:Menthol reductase (MMR) ]] |
| − | rect | + | |
| − | rect | + | rect 929 67 1018 85 [[CPDMO]] |
| − | rect | + | rect 700 10 759 30 [[Menthofuran synthase (MFS)]] |
| − | rect | + | |
| − | rect | + | rect 704 283 759 304 [[ Limonene-3-hydroxylase (L3H)]] |
| − | rect | + | rect 699 213 755 233 [[ Isopiperitenol Dehydrogenase (IPDH)]] |
| − | rect | + | rect 701 137 756 157 [[ Isopiperitenol Reductase (IPR)]] |
| − | rect | + | rect 700 57 756 77 [[Isopulegone Isomerase (IPGI)]] |
| − | rect | + | rect 355 212 414 233 [[HDS]] |
| − | rect | + | rect 469 213 525 231 [[Hydroxymethylbutenyl diphosphate reductase (HMBPPR)]] |
| − | rect | + | |
| − | rect | + | rect 705 363 760 383 [[Limonene-6-Hydroxylase (L6H)]] |
| + | rect 703 436 760 455 [[ Carveol Dehydrogenase (CDH)]] | ||
| + | rect 704 512 760 532 [[OYE]] | ||
| + | rect 699 587 767 605 [[CHMO]] | ||
| + | |||
| + | rect 24 72 81 93 [[DXS]] | ||
| + | rect 20 135 77 154 [[DXR]] | ||
| + | rect 16 212 71 230 [[CMS]] | ||
| + | rect 119 212 172 231 [[CMK]] | ||
| + | rect 239 210 294 231 [[MCS]] | ||
| + | rect 585 205 641 226 [[IDI]] | ||
| + | |||
| + | rect 149 26 208 44 [[AACT]] | ||
| + | rect 148 99 216 119 [[HMGS]] | ||
| + | rect 226 136 291 155 [[HMGR]] | ||
desc bottom-right | desc bottom-right | ||
Revision as of 10:55, 12 August 2016
This wiki page describes the construction and simulation of a kinetic model of Monoterpenoid Biosynthesis.
Contents
Monoterpenoid Biosynthesis
Description of the model
To find out how each enzyme in the network is modelled, click on the rectangles in the figure on the left. Alternatively, you can click the enzyme names from the list below.
IPP and DMAPP Biosynthesis
Diphosphomelvalonate decarboxylase (DMD)
Hydroxymethylbutenyl diphosphate reductase (HMBPPR)
Limonene Biosynthesis
Geranyl diphosphate synthase (GPPS)
Peppermint Biosynthesis
Isopiperitenol Dehydrogenase (IPDH)
Isopiperitenol Reductase (IPR)
Mint Biosynthesis
Menthone:Neomenthol reductase (MNMR)
Menthone:Menthol reductase (MMR)
Spearmint Biosynthesis
Methanofuran Biosynthesis
Reversible Michaelis-Menten equation
All reactions in this model are described using reversible Michaelis-Menten equation.
Kinetic Parameters
Strategies for Estimating Kinetic Parameter Values
Using Equilibrium Constant (Keq) in the reversible Michaelis-Menten equation
Using the equilibrium constant in the reversible Michaelis-Menten reaction reduces the need to obtain or estimate Vmaxreverse parameter value, which is often not available in literature.
Using the Haldane relationship, the equilibrium constant (Keq) can be written as:
- Failed to parse (Cannot store math image on filesystem.): {\color{Red}K_\mathrm{eq}} = \frac{Vmax_\mathrm{forward} * Km_\mathrm{product} }{Vmax_\mathrm{reverse} * Km_\mathrm{substrate}}
The reversible Michaelis-Menten rate equation can then be rewritten to contain the equilibrium constant (Keq) instead of using the ratio of the Vmax values.
- Failed to parse (Cannot store math image on filesystem.): v_\mathrm{reaction} = Vmax_\mathrm{forward} * \cfrac {\cfrac{[S]}{Km_\mathrm{substrate}} * \left ( 1 - \cfrac {[P]}{[S]* \color{red}K_\mathrm{eq}} \right )} {1 + \cfrac {[S]}{Km_\mathrm{substrate}} + \cfrac {[P]}{Km_\mathrm{product}}}
where [S] and [P] corresponds to the concentration of the substrate and product respectively.
Calculating the Equilibrium Constant (Keq)
The equilibrium constant can be calculated from the ratio of the forward and reverse reaction rates as described in the above section, given that the values for these rates are known. Unfortunately, the information for these values are often difficult to find, especially for reverse reaction rates (Vmaxreverse).
As an alternative, the equlibrium constant, Keq, can also be calculated from the Gibbs free energy of a reaction, ΔGr, using the Van't Hoff isotherm equation:
and by dividing both sides of the equation with RT, and later take the exponents of both sides, the Keq can be calculated by this equation:
where;
| Keq | Equilibrium constant |
| -ΔG° | Gibbs free energy change |
| R | Gas constant with a value of 8.31 JK-1mol-1 |
| T | Temperature which is always expressed in kelvin |
List of kinetic parameter values
Lists of values retrieved from published and unpublished sources for the kinetics of all the enzymes included in this study can be found here.
Data gathering progress:
Legend:
| Have not started ·· | 1 -2 data found ·· | 3-4 data found ·· | sufficient data found/estimated ·· | data distribution generated ·· | data sampled |
| to do | DONE! |
| Enzyme | Pathway | Keq | Kcat | VmaxF | KmSub1 | KmSub2 | [Sub1] | [Sub2] | VmaxR | KmProd1 | KmProd2 | [Prod1] | [Prod2] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01_GPPS | Limonene Biosynthesis | 1 | 3 | 4 | 4 | 4 | 3 | 3 | |||||
| 01_L3H | Peppermint Biosynthesis | 1 | 1 | 1 | 1 | ||||||||
| 01_L6H | Spearmint Biosynthesis | 1 | 1 | 1 | 1 | 1 | |||||||
| 01_MFS | Methanofuran Biosynthesis | ||||||||||||
| 01_PGR | Mint Biosynthesis | ||||||||||||
| 02_CDH | Spearmint Biosynthesis | ||||||||||||
| 02_IPDH | Peppermint Biosynthesis | ||||||||||||
| 02_LimSynth | Limonene Biosynthesis | 1 | 5 | 3 | >6 | no Sub2 | 4 | no Sub2 | estimated | estimated | estimated | estimated | estimated |
| 02_MNMR (menthone) | Mint Biosynthesis | 1 | 1 | ||||||||||
| 02a_IPDH | Spearmint Biosynthesis | ||||||||||||
| 03_IPR | Peppermint Biosynthesis | ||||||||||||
| 03_MMR (menthone) | Mint Biosynthesis | 1 | 1 | ||||||||||
| 03_OYE | Spearmint Biosynthesis | ||||||||||||
| 04_CHMO | Spearmint Biosynthesis | ||||||||||||
| 04_IPGI | Peppermint Biosynthesis | ||||||||||||
| 04_MNMR (isomenthone) | Mint Biosynthesis | 1 | 1 | ||||||||||
| 04_MNMR (isomenthone) | Mint Biosynthesis | 1 | 1 |
Abbreviations
List of abbreviations used in this study can be found here
Back to the main model page.