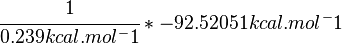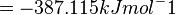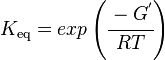Limonene-6-Hydroxylase (L6H)
You can go back to main page of the kinetic model here.
Contents
What we know
This enzyme is also known as (S)-limonene 6-monooxygenase, (-)-limonene 6-hydroxylase, (-)-limonene 6-monooxygenase, (-)-limonene,NADPH:oxygen oxidoreductase (6-hydroxylating)
Reaction catalysed
Metabolite and Enzyme Background Information
Long metabolite and enzyme names are abbreviated in the model for clarity and standard identification purposes.
| Metabolite | Abbreviation | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
|---|---|---|---|---|---|---|---|
| (-)-4S-limonene | Limonene | C10H16 | 136.24 | 15384 | 449062 | 22311 or 439250 | |
| (-)-trans-carveol | carveol | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
| NADPH | NADPH | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
| NADP | NADP | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
| Limonene-6-hydroxylase | L6H | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | LIMONENE-6-MONOOXYGENASE-RXN |
| Metabolite | Abbreviation | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
Equation Rate
This reaction is modelled using the reversible Michaelis-Menten equation, with two substrates; Limonene and NADPH, and two products; Carveol and NADP.
| Parameter | Description | Units |
|---|---|---|
| VL6H | Net reaction rate for Limonene-6-Hydroxylase | Unit |
| Vmaxforward | Maximum reaction rate towards the production of trans-carveol | Unit |
| Kmlimonene | Michaelis-Menten constant for Limonene | Unit |
| Kmcarveol | Michaelis-Menten constant for trans-carveol | Unit |
| KmNADPH | Michaelis-Menten constant for NADPH | Unit |
| KmNADP | Michaelis-Menten constant for NADP | Unit |
| Keq | Equilibrium constant | Unit |
| [Limonene] | Limonene concentration | Unit |
| [Carveol] | trans-carveol concentration | Unit |
| [NADPH] | NADPH concentration | Unit |
| [NADP] | NADP concentration | Unit |
Strategies for estimating the kinetic parameter values
Calculating the Equilibrium Constant
The equilibrium constant can be calculated using the Van't Hoff Isotherm equation, which requires the information on the enzyme's standard Gibbs free energy [1].
Standard Gibbs Free energy
Standard Gibbs free energy is -92.52051 kcal·mol-1 [2] according to MetaCyc [[1]].
SI derived unit for Gibbs free energy is Joules per mol (J mol-1). 1 kJ·mol−1 is equal to 0.239 kcal·mol−1.
Therefore, the Gibbs free energy for L6H in kJ mol-1 is:
The equilibrium constant
Using the Van't Hoff Isotherm equation:
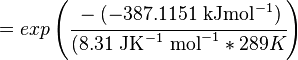
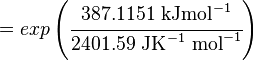

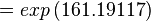
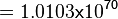
where;
| Keq | Equilibrium constant |
| -∆G° | Gibbs free energy change. For L6H it is 387.1151 kJmol-1 |
| R | Gas constant with a value of 8.31 JK-1mol-1 |
| T | Temperature which is always expressed in kelvin |
Published Kinetic Parameter Values
Km Values
| Km (mM) | Unit | Substrate / Product | Directionality | Organism | References |
|---|---|---|---|---|---|
| 20 | µM | Carveol | forward | Mentha sp. | [3] |
| value | µM | susbstrate | directionality | organism | ref |
| value | µM | susbstrate | directionality | organism | ref |
| value | µM | susbstrate | directionality | organism | ref |
Vmax values
| Vmax | Unit | Directionality | Organism | References |
|---|---|---|---|---|
| 64 | nmol/mg.h | forward | Mentha sp. | [3] |
| val | µmol/min/mg (unit) | directionality | Organism | References |
Kcat values
| Kcat | Unit | Organism | Reference |
|---|---|---|---|
| value | s-1 | Organism | ref e.g. Alonso 1992 [4] |
Extracting Information from (INSERT SUBSTRATE/PRODUCT) Production Rates
| Amount produced (mg/L) | Time (H) | Organism | Description | Reaction Flux (µM/s) |
|---|---|---|---|---|
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
Published Kinetic Parameter Values
| Km (mM) | Vmax | Kcat (s-1) | Kcat/Km | Organism | Description |
|---|---|---|---|---|---|
| 0.00125 | - | - | Z | A -> B | |
| 0.0018 | - | - | - | Z | A -> B |
| Y | Y | Y | Y | Z | A -> B |
| Y | Y | Y | Y | Z | A -> B |
| Y | Y | - | - | Z | A -> B |
| Y | - | Y | - | Z | GPP -> B |
| Y | - | Y | - | Z | GPP -> B |
| x | - | y | - | Z. | A -> B |
Detailed descriptions of kinetic values used in this model
A more detailed description of the kinetic values listed above can be found here.
Simulations
References
- ↑ Liebermeister, W. & Klipp, E. 2005. http://pubman.mpdl.mpg.de/pubman/item/escidoc:1585440/component/escidoc:1585439/Liebermeister+et+al.+-+IEE+Proc.-Syst.+Biol.pdf."Biochemical networks with uncertain parameters." IEE Proc-Syst. Biol. 152(3): 97-105
- ↑ Latendresse, M. 2013. http://www.biocyc.org/PGDBConceptsGuide.shtml#gibbs. "Computing Gibbs Free Energy of Compounds and Reactions in MetaCyc."
- ↑ 3.0 3.1 Karp, F. et. al. 1990. "Monoterpene biosynthesis: Specificity of the hydroxylations of (-)-Limonene by enzyme preparations from Peppermint (Mentha piperita), Spearmint (Mentha spicata), and Perilla (Perilla frutescens) leaves", Archives of Biochemistry and Biophysics, 276(1):219-226
- ↑ Alonso et. al. 1992. "Purification of 4S-Limonene Synthase, a Monoterpene Cyclase from the Glandular Trichomes of Peppermint (Mentha x piperita) and Spearmint (Mentha spicata)", The Journal of Biological Chemistry, 267(11):7582-7587
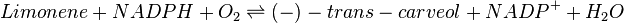
![V_\mathrm{L6H} = Vmax_\mathrm{forward} * \cfrac {\cfrac{[Limonene]}{Km_\mathrm{Limonene}} * \left ( 1 - \cfrac {[Limonene]*[PP]}{[GPP]*K_\mathrm{eq}} \right )}{1 + \cfrac {[GPP]}{Km_\mathrm{GPP}} + \cfrac {[Limonene]}{Km_\mathrm{Limonene}} + \cfrac {[PP]}{Km_\mathrm{PP}} + \cfrac {[Limonene]*[PP]}{Km_\mathrm{Limonene}*Km_\mathrm{PP}}}](/wiki/images/math/6/e/6/6e68d78babd5acdaa586f13c500e95dd.png)