Difference between revisions of "Kinetic model of Central Metabolism"
(→Quantifying the flux) |
(→Quantifying the flux) |
||
| Line 113: | Line 113: | ||
<math>\text{Fraction Unlabeled} = \frac{\text{Peak Height}_{\text{unlabeled form}}}{\text{Sum of peak height}_{\text{every isotopic form}}}</math> | <math>\text{Fraction Unlabeled} = \frac{\text{Peak Height}_{\text{unlabeled form}}}{\text{Sum of peak height}_{\text{every isotopic form}}}</math> | ||
| − | * If we can calculate the apparent first order rate constant <math>K_X</math> then the flux can be calculated as <math>f_X = K_X \times X^T</math> | + | * If we can calculate the apparent first order rate constant <math>K_X</math> then the flux can be calculated as <math>f_X = K_X \times X^T</math>. [<math>X^T</math> is the intracellular concentration of X]. |
| + | |||
| + | * With <math>X^T</math> at steady-state, we have | ||
| + | <math>\frac{dX^{U}}{dt} = -f_X(X^{U}/X^{T}</math> | ||
| + | |||
| + | The analytical solution is | ||
| + | <math>\frac{X^U}{X^T} = exp(-f_Xt/X^T)</math> | ||
==References== | ==References== | ||
<references/> | <references/> | ||
Revision as of 11:23, 11 November 2014
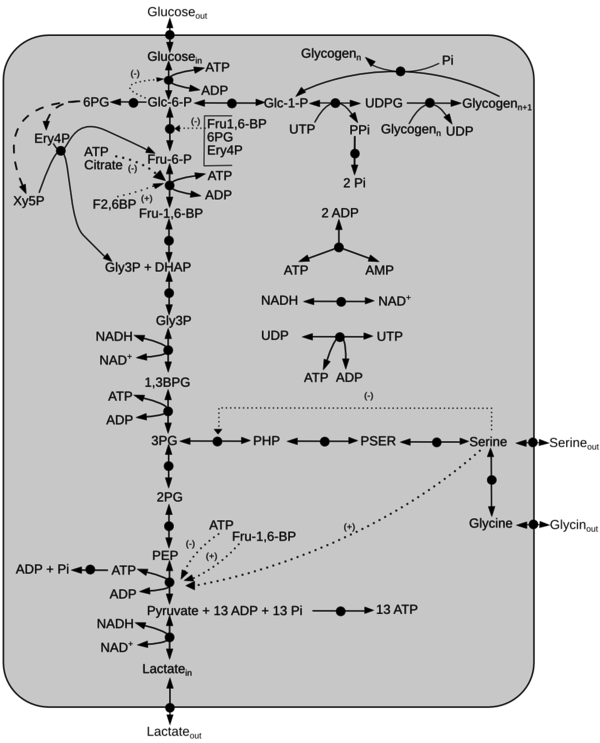
A kinetic model of glycolysis with serine activation is constructed from the literature data [1][2][3][4][5].
Contents
Description of the model
Schematic diagram of the model is given here. The dotted line represents activation(+) or inhibition(-) and the dashed arrow indicate Pentose Phosphate Pathway reactions not included in the model. Click on a reaction to have more information
Reactions
Details of the abbreviations for this model is listed here. Reactions of the model are listed below.
Initial concentration of the metabolites can be found here
Model File
The SBML file of the model can be found here
Notes
Necessary observation during model building and simulation is noted here.
Global parameters
Calculating 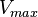
Some of the Vmax value in the paper "Modeling cancer glycolysis" is given in 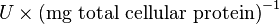 unit [1]. To homogenize the units it is then converted back to
unit [1]. To homogenize the units it is then converted back to 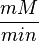 by multiplying with 65 as the HeLa cell was incubated in
by multiplying with 65 as the HeLa cell was incubated in 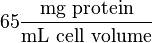 .
.
- Converting
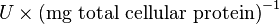 to
to 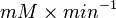 :
:
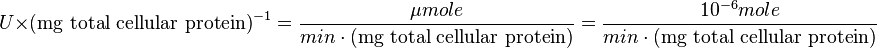
Also, 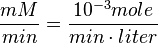
We want to convert 1 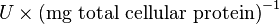 to
to 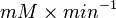 where HeLa cell was incubated in
where HeLa cell was incubated in 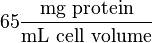
Therefore, 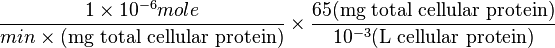
=> 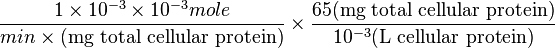
Striking out the necessary elements we get
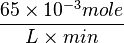
=> 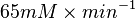
Quantifying the flux
We used the Kinetic Flux Profiling method to quantify the flux values.
- First we determine the unlabeled from for each metabolites
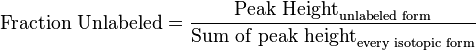
- If we can calculate the apparent first order rate constant
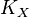 then the flux can be calculated as
then the flux can be calculated as 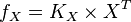 . [
. [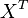 is the intracellular concentration of X].
is the intracellular concentration of X].
- With
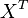 at steady-state, we have
at steady-state, we have
Failed to parse (Cannot store math image on filesystem.): \frac{dX^{U}}{dt} = -f_X(X^{U}/X^{T}
The analytical solution is
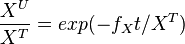
References
- ↑ 1.0 1.1 Marín-Hernández A, Gallardo-Pérez JC, Rodríguez-Enríquez S et al (2011). Modeling cancer glycolysis. Biochim Biophys Acta, 1807:755–767 (doi)
- ↑ Turnaev II, Ibragimova SS, Usuda Y et al (2006). Mathematical modeling of serine and glycine synthesis regulation in Escherichia coli. Proceedings of the fifth international conference on bioinformatics of genome regulation and structure 2:78–83
- ↑ Smallbone K, Stanford NJ (2013). Kinetic modeling of metabolic pathways: Application to serine biosynthesis. In: Systems Metabolic Engineering, Humana Press. pp. 113–121
- ↑ Palm, D.C. (2013). The regulatory design of glycogen metabolism in mammalian skeletal muscle (Ph.D.). University of Stellenbosch
- ↑ Ettore Murabito (2010). Application of differential metabolic control analysis to identify new targets in cancer treatment (Ph.D.). University of Manchester