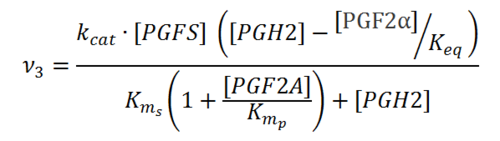Difference between revisions of "Transformation of PGH2 to PGF2α"
(Created page with "== Reaction == ==Chemical equation== <center><math> FA \rightleftharpoons AA </math></center> == Rate equation == == Parameters == {|class="wikitable" ! Name ! Value ! ...") |
|||
| (28 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
| + | [[Welcome to the In-Silico Model of Cutaneous Lipids Wiki | Return to overview]] | ||
| + | |||
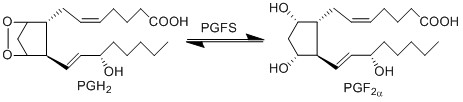
| + | The isomerisation of PGH2 to PGF2α is performed by PGFS. This reaction results in the formation of two hydroxyl groups at C9 and C11. Two isoforms of PGFS have been reported in the literature in humans, the aldo-keto reductase family 1-member C3 isoform (encoded by the AKR1C3 gene) and the prostamide/prostaglandin F synthase isoform (encoded by the PRXL2B gene). | ||
| + | |||
| + | |||
== Reaction == | == Reaction == | ||
| + | |||
| + | [[File:R3_PGH2_-PGF2A.jpg|center|500px]] | ||
==Chemical equation== | ==Chemical equation== | ||
| − | <center><math> | + | <center><math> PGH2 \rightleftharpoons PGF2a </math></center> |
== Rate equation == | == Rate equation == | ||
| − | == Parameters == | + | [[File:R03.PNG|center|500px]] |
| + | |||
| + | == Enzyme Parameters == | ||
| + | |||
| + | === K<sub>ms</sub> === | ||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |1.90E-03 ± 1.5E-03 | ||
| + | |<math> mM </math> | ||
| + | |Human | ||
| + | |Placental Aldose Reductase (AKR1B1), E. Coli, pH 7, 37°C | ||
| + | |512 | ||
| + | |<ref name="Kabututu2009"> [http://www.ncbi.nlm.nih.gov/pubmed/19010934 Z. Kabututu, M. Manin, Prostaglandin F2alpha synthase activities of aldo-keto reductase 1B1, 1B3 and 1B7.'' J Biochem 2009,145(2):161-8)]</ref> | ||
| + | |- | ||
| + | |1.80E-02 | ||
| + | |<math> mM </math> | ||
| + | |Human | ||
| + | |Lung PGF2a Synthase (AKR1C3), E. Coli, pH 7, 37°C | ||
| + | | 512 | ||
| + | |<ref name="Kabututu2009"> [http://www.ncbi.nlm.nih.gov/pubmed/19010934 Z. Kabututu, M. Manin, Prostaglandin F2alpha synthase activities of aldo-keto reductase 1B1, 1B3 and 1B7.'' J Biochem 2009,145(2):161-8)]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGFS Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.33E-02 || 3.81E+00 || -3.45E+00 || 9.33E-01 | ||
| + | |} | ||
| + | |||
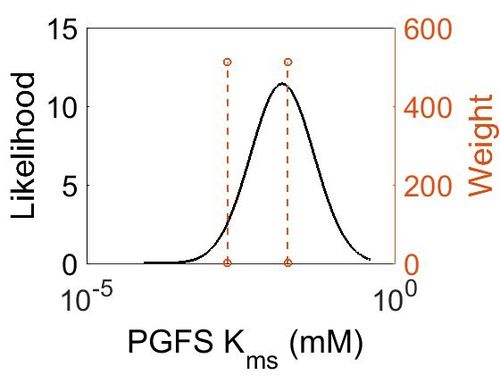
| + | [[Image:9_(2).jpg|none|thumb|500px|The estimated probability distribution for PGFS Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === K<sub>mp</sub> === | ||
| + | This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail[[Quantification of parameter uncertainty | here]]). As a result, no confidence interval factor or literature values were cited for this parameter. | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGFS Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.46E-02 || -2.86E+00 || 1.17E+00 | ||
| + | |} | ||
| + | |||
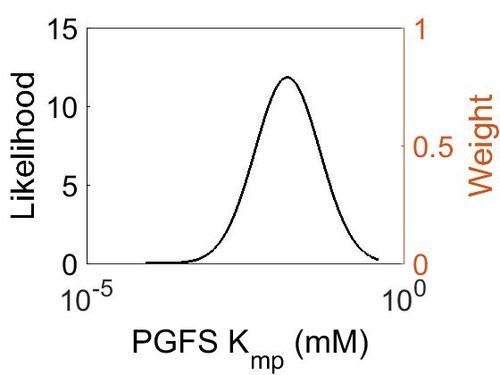
| + | [[Image:10_(2).jpg|none|thumb|500px|The estimated probability distribution for PGFS Kmp. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| − | {|class="wikitable" | + | === k<sub>cat</sub> === |
| − | + | {|class="wikitable sortable" | |
| + | |+ style="text-align: left;" | Literature values | ||
| + | |- | ||
! Value | ! Value | ||
! Units | ! Units | ||
! Species | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| − | | | + | | 14.9 |
| − | | | + | | per minute |
| − | | | + | | Mouse/Swine |
| − | | | + | | Prostamide/PGF Synthase Purified |
| − | | | + | from Swine Brain and of Murine Enzyme Expressed in E. coli |
| + | |128 | ||
| + | | <ref name="Moriuchi2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18006499 Moriuchi H., Molecular characterization of a novel type of prostamide/prostaglandin F synthase, belonging to the thioredoxin-like superfamily.'' J Biol Chem. 2008 Jan 11;283(2):792-801.)]</ref> | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGFS kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.48E+01 || 1.10E+00 || 2.71E+00 || 9.49E-02 | ||
|} | |} | ||
| + | |||
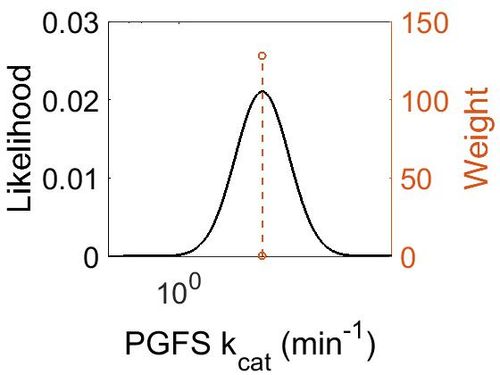
| + | [[Image:11.jpg|none|thumb|500px|The estimated probability distribution for PGFS kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === Enzyme concentration === | ||
| + | |||
| + | To convert the enzyme concentration from ppm to mM, the following [[Common equations#Enzyme concentration (mM)|equation]] was used. | ||
| + | |||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |105 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Platelet | ||
| + | Enzyme: PGFS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
| + | |- | ||
| + | |19.9 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Lung | ||
| + | Enzyme: PGFS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |40.3 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Esophagus | ||
| + | Enzyme: PGFS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |110 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Oral Cavity | ||
| + | Enzyme: PGFS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGFS concentration distribution | ||
| + | ! Mode (ppm) !! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 6.41E+01 || 3.55E-04 || 2.05E+00 || 4.53E+00 || 6.06E-01 | ||
| + | |} | ||
| + | |||
| + | |||
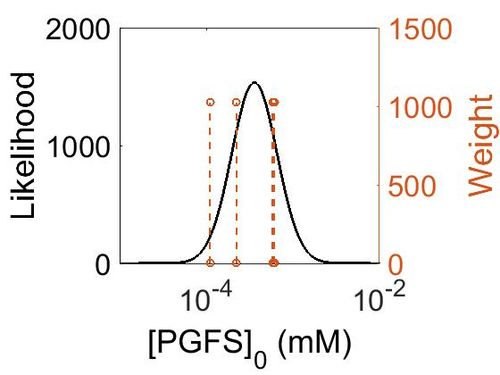
| + | [[Image:163.jpg|none|thumb|500px|The estimated probability distribution for PGFS concentration. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === K<sub>eq</sub> === | ||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Gibbs Free Energy Change | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |(-6.64) | ||
| + | |kcal/mol | ||
| + | |Not stated | ||
| + | |Estimated | ||
| + | Enzyme: PGFS | ||
| + | Substrate: Arachidonate | ||
| + | Product: PGF2a | ||
| + | pH: 7.3 | ||
| + | ionic strength: 0.25 | ||
| + | |64 | ||
| + | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=PROSTAGLANDIN-E-SYNTHASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
| + | |||
| + | |} | ||
| + | {| class="wikitable" | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 7.46E+04 || 1.00E+01 || 1.20E+01 || 8.90E-01 | ||
| + | |} | ||
| + | |||
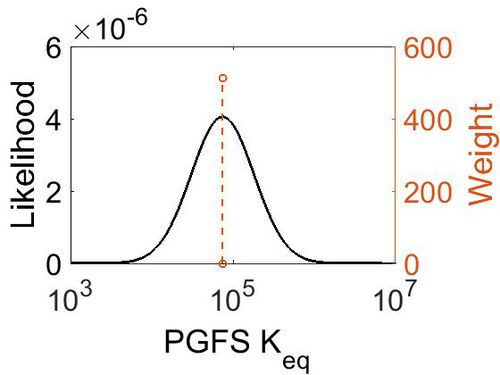
| + | [[Image:12.jpg|none|thumb|500px|The estimated probability distribution for PGFS Keq. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | == References == | ||
| + | <references/> | ||
| + | |||
| + | |||
| + | == Related Reactions == | ||
| + | * [[Transformation of AA to PGH2 |Transformation of AA to PGH2]] | ||
Latest revision as of 08:36, 21 August 2019
The isomerisation of PGH2 to PGF2α is performed by PGFS. This reaction results in the formation of two hydroxyl groups at C9 and C11. Two isoforms of PGFS have been reported in the literature in humans, the aldo-keto reductase family 1-member C3 isoform (encoded by the AKR1C3 gene) and the prostamide/prostaglandin F synthase isoform (encoded by the PRXL2B gene).
Contents
Reaction
Chemical equation
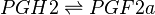
Rate equation
Enzyme Parameters
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 1.90E-03 ± 1.5E-03 | 
|
Human | Placental Aldose Reductase (AKR1B1), E. Coli, pH 7, 37°C | 512 | [1] |
| 1.80E-02 | 
|
Human | Lung PGF2a Synthase (AKR1C3), E. Coli, pH 7, 37°C | 512 | [1] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.33E-02 | 3.81E+00 | -3.45E+00 | 9.33E-01 |
Kmp
This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail here). As a result, no confidence interval factor or literature values were cited for this parameter.
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 1.46E-02 | -2.86E+00 | 1.17E+00 |
kcat
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 14.9 | per minute | Mouse/Swine | Prostamide/PGF Synthase Purified
from Swine Brain and of Murine Enzyme Expressed in E. coli |
128 | [2] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.48E+01 | 1.10E+00 | 2.71E+00 | 9.49E-02 |
Enzyme concentration
To convert the enzyme concentration from ppm to mM, the following equation was used.
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 105 | 
|
Human | Expression Vector: Platelet
Enzyme: PGFS pH: 7.5 Temperature: 37 °C |
1024 | [3] |
| 19.9 | 
|
Human | Expression Vector: Lung
Enzyme: PGFS pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| 40.3 | 
|
Human | Expression Vector: Esophagus
Enzyme: PGFS pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| 110 | 
|
Human | Expression Vector: Oral Cavity
Enzyme: PGFS pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| Mode (ppm) | Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|---|
| 6.41E+01 | 3.55E-04 | 2.05E+00 | 4.53E+00 | 6.06E-01 |
Keq
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| (-6.64) | kcal/mol | Not stated | Estimated
Enzyme: PGFS Substrate: Arachidonate Product: PGF2a pH: 7.3 ionic strength: 0.25 |
64 | [5] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 7.46E+04 | 1.00E+01 | 1.20E+01 | 8.90E-01 |
References
- ↑ 1.0 1.1 Z. Kabututu, M. Manin, Prostaglandin F2alpha synthase activities of aldo-keto reductase 1B1, 1B3 and 1B7. J Biochem 2009,145(2):161-8)
- ↑ Moriuchi H., Molecular characterization of a novel type of prostamide/prostaglandin F synthase, belonging to the thioredoxin-like superfamily. J Biol Chem. 2008 Jan 11;283(2):792-801.)
- ↑ M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ 4.0 4.1 4.2 M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471