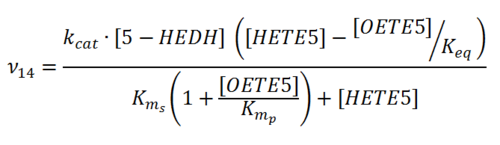Difference between revisions of "Transformation of 5-HETE to 5-OXO-ETE"
(→Parameters) |
|||
| Line 2: | Line 2: | ||
The metabolism of 5-HETE to 5-Oxo-ETE is catalysed by the enzyme 5-Hydroxyeicosanoid dehydrogenase (5-HEDH). This enzyme acts reversibly and has been reported to be found in eosinophils, monocytes, DC, B-lymphocytes, keratinocytes and platelets (Steinhilber book) *skin (need to check). The product binds with the OXE receptor (Hosoi2002) and is 30-100 more bioactive than 5-HETE. 5-Oxo-ETE is a chemoattractant for neutrophils and eosinophils. | The metabolism of 5-HETE to 5-Oxo-ETE is catalysed by the enzyme 5-Hydroxyeicosanoid dehydrogenase (5-HEDH). This enzyme acts reversibly and has been reported to be found in eosinophils, monocytes, DC, B-lymphocytes, keratinocytes and platelets (Steinhilber book) *skin (need to check). The product binds with the OXE receptor (Hosoi2002) and is 30-100 more bioactive than 5-HETE. 5-Oxo-ETE is a chemoattractant for neutrophils and eosinophils. | ||
| − | |||
| − | |||
== Reaction == | == Reaction == | ||
| Line 19: | Line 17: | ||
== Parameters == | == Parameters == | ||
| − | Note | + | Note that there was limited availability of kinetic data for this specific enzyme, therefore parameters for general alcohol dehydrogenase (ADH) was used when non were available. |
| + | === K<sub>ms</sub> === | ||
{|class="wikitable" | {|class="wikitable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature Values |
! Value | ! Value | ||
! Units | ! Units | ||
| Line 62: | Line 61: | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the 5-HEDH Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 6.34E-04 || 1.63E+01 || -6.31E+00 || 1.03E+00 | ||
| + | |} | ||
| + | |||
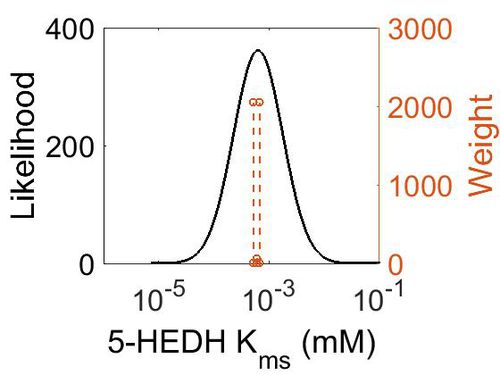
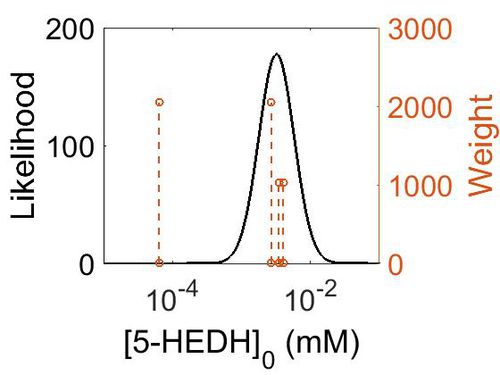
| + | [[Image:45.jpg|none|thumb|500px|The estimated probability distribution for 5-HEDH Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===K<sub>mp</sub> === | ||
| + | This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail[[Quantification of parameter uncertainty | here]]). As a result, no confidence interval factor or literature values were cited for this parameter. | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the 5-HEDH Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.14E+04 || 1.09E+01 || 1.25E+00 | ||
| + | |} | ||
| + | |||
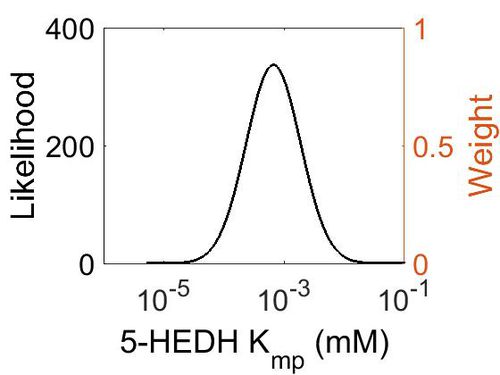
| + | [[Image:46.jpg|none|thumb|500px|The estimated probability distribution for 5-HEDH Kmp. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===V<sub>max</sub> === | ||
{|class="wikitable" | {|class="wikitable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
! Value | ! Value | ||
! Units | ! Units | ||
| Line 98: | Line 121: | ||
|} | |} | ||
| + | === k<sub>cat</sub> === | ||
{|class="wikitable" | {|class="wikitable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
! Value | ! Value | ||
! Units | ! Units | ||
| Line 120: | Line 144: | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the 5-HEDH kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.15E+04 || 4.02E+01 || 1.09E+01 || 1.25E+00 | ||
| + | |} | ||
| + | |||
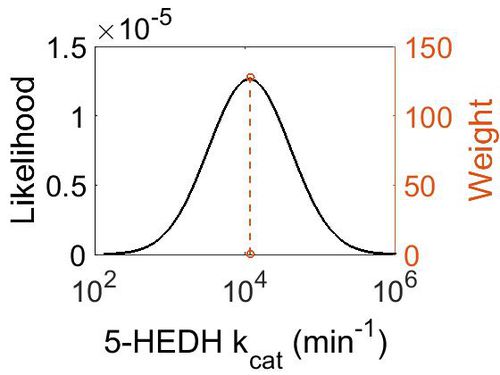
| + | [[Image:47.jpg|none|thumb|500px|The estimated probability distribution for 5-HEDH kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | === Enzyme concentration (Using ADH5 abundance data) === | ||
{|class="wikitable" | {|class="wikitable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
! Value | ! Value | ||
! Units | ! Units | ||
| Line 167: | Line 202: | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the 5-HEDH concentration distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 4.88E+02 || 6.37E+00 || 7.49E+00 || 1.14E+00 | ||
| + | |} | ||
| + | |||
| + | [[Image:155.jpg|none|thumb|500px|The estimated probability distribution for 5-HEDH concentration. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===K<sub>eq</sub>=== | ||
{|class="wikitable" | {|class="wikitable" | ||
| − | |+ style="text-align: left;" | Gibbs Free energy | + | |+ style="text-align: left;" | Literature values |
| − | + | ! Gibbs Free energy | |
! Units | ! Units | ||
! Species | ! Species | ||
| Line 189: | Line 236: | ||
|<ref name="MetaCyc”>[https://metacyc.org/META/NEW-IMAGE?type=REACTION&object=RXN-16416 Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[https://metacyc.org/META/NEW-IMAGE?type=REACTION&object=RXN-16416 Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the 5-HEDH Keq distribution | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.12E-01 || 1.00E+01 || -1.39E+00 || 8.91E-01 | ||
| + | |} | ||
| + | |||
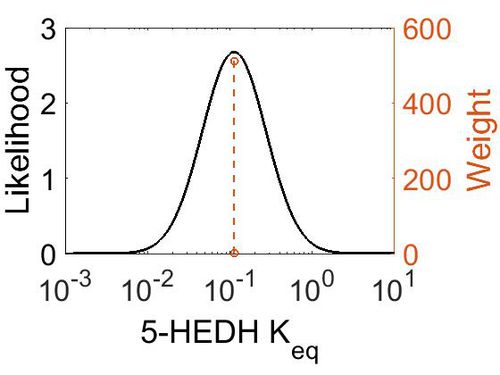
| + | [[Image:48.jpg|none|thumb|500px|The estimated probability distribution for 5-HEDH Keq. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
== Related Reactions == | == Related Reactions == | ||
Revision as of 12:21, 15 May 2019
The metabolism of 5-HETE to 5-Oxo-ETE is catalysed by the enzyme 5-Hydroxyeicosanoid dehydrogenase (5-HEDH). This enzyme acts reversibly and has been reported to be found in eosinophils, monocytes, DC, B-lymphocytes, keratinocytes and platelets (Steinhilber book) *skin (need to check). The product binds with the OXE receptor (Hosoi2002) and is 30-100 more bioactive than 5-HETE. 5-Oxo-ETE is a chemoattractant for neutrophils and eosinophils.
Contents
Reaction
Chemical equation
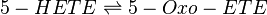
Rate equation
Parameters
Note that there was limited availability of kinetic data for this specific enzyme, therefore parameters for general alcohol dehydrogenase (ADH) was used when non were available.
Kms
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 0.0006 | mM | Unkown | Follows a ping-pong mechanism by binding with NADP+, transforming it to NADPH and releasing it before binding 5S-HETE. | [1] |
| 0.00067 | mM | Human Cell lines | Follows a ping-pong mechanism by binding with NADP+ (Km 139 nM), transforming it to NADPH (inhibited by NADPH (Ki 224 nM)) and releasing it before binding 5S-HETE.
Method: In vitro Organism: Human Expression vector: U937 and HL-60 cell lines Enzyme:5-HEDH pH: 7.4 Temperature: 37 ◦C |
[2] |
| 0.000516 ± 0.00019 | mM | Human Cell lines |
Method: In vitro Organism: Human Expression vector: U937 Enzyme:5-HEDH pH: 7.4 Temperature: 37 ◦C |
[3] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 6.34E-04 | 1.63E+01 | -6.31E+00 | 1.03E+00 |
Kmp
This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail here). As a result, no confidence interval factor or literature values were cited for this parameter.
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 1.14E+04 | 1.09E+01 | 1.25E+00 |
Vmax
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 0.54 ± 0.30 | pmol/ mi mg | Human Cell lines - Neutrophils |
Method: In vitro Organism: Human Expression vector: U937 and HL-60 cell lines Enzyme:5-HEDH pH: 7.4 Temperature: 37 ◦C |
[2] |
| 0.40 ± 0.12 | pmol/ mi mg | Human Cell lines - U937 |
Method: In vitro Organism: Human Expression vector: U937 and HL-60 cell lines Enzyme:5-HEDH pH: 7.4 Temperature: 37 ◦C |
[2] |
kcat
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 11640 ± 840 | min-1 | Yokenella | Method: In vitro
Organism: Human Expression vector: E Coli Enzyme: pH: 6.5 Temperature: 65 ◦C Substrate: NADPH |
B |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.15E+04 | 4.02E+01 | 1.09E+01 | 1.25E+00 |
Enzyme concentration (Using ADH5 abundance data)
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 490 | ppm | Human | Expression Vector: Skin
Enzyme: ADH5 pH: 7.5 Temperature: 37 °C |
[4] |
| 625 | ppm | Human | Expression Vector: Oral Cavity
Enzyme: ADH5 pH: 7.5 Temperature: 37 °C |
[4] |
| 728 | ppm | Human | Expression Vector: Esophagus
Enzyme: ADH5 pH: 7.5 Temperature: 37 °C |
[5] |
| 11.5 | ppm | Human | Expression Vector: Skin
Enzyme: ADH5 pH: Unknown Temperature: Unknown |
[5] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 4.88E+02 | 6.37E+00 | 7.49E+00 | 1.14E+00 |
Keq
| Gibbs Free energy | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 1.29 | kcal/mol | Arabidopsis thaliana col | Same reaction, no prediction available for 5-HEDH on Metacyc
Estimated Enzyme: ω-hydroxy fatty acid ω-alcohol dehydrogenase Substrate: 18-hydroxyoleate Product: 18-oxo-oleate pH: 7.3 ionic strength: 0.25 |
[6] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.12E-01 | 1.00E+01 | -1.39E+00 | 8.91E-01 |
Related Reactions
References
- ↑ Lipoxygenases in Inflammation - edited by Dieter Steinhilber
- ↑ 2.0 2.1 2.2 [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1828885/pdf/bj4030157.pdf Karl-Rudolf ERLEMANN, Regulation of 5-hydroxyeicosanoid dehydrogenase activity in monocytic cells, Biochem. J. (2007) 403, 157–165]
- ↑ [http://jpet.aspetjournals.org/content/jpet/329/1/335.full.pdf Karl-Pranav Patel, Selectivity of 5-Hydroxyeicosanoid Dehydrogenase and Its Inhibition by 5-Hydroxy-Long-Chain Fatty Acids, Journal of Pharmacology and Experimental Therapeutics April 2009, 329 (1) 335-341; ]
- ↑ 4.0 4.1 [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587]
- ↑ 5.0 5.1 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471