Difference between revisions of "Formation of homo-dimer R2"
(→Parameters with uncertainty) |
(→Parameters with uncertainty) |
||
| Line 53: | Line 53: | ||
More specifically, the weight of the sampling is kept at <math> 30 nM </math> (value that corresponds to the wild type protein homodimerization) which is set as the mode of the log-normal distribution for the <math>K_{d6}</math>. However, we wish to explore the full nanomolar scale when sampling for parameter values and therefore the confidence interval factor is set to <math> 30 </math>. In this way, the range where 95.45% of the values are found is between <math>1</math> and <math>900 nM </math>. | More specifically, the weight of the sampling is kept at <math> 30 nM </math> (value that corresponds to the wild type protein homodimerization) which is set as the mode of the log-normal distribution for the <math>K_{d6}</math>. However, we wish to explore the full nanomolar scale when sampling for parameter values and therefore the confidence interval factor is set to <math> 30 </math>. In this way, the range where 95.45% of the values are found is between <math>1</math> and <math>900 nM </math>. | ||
| − | With regards to the parameter <math>k_{on6}</math>, in order to explore the full range of plausible values, the mode of the log-normal distribution is set to <math>0. | + | With regards to the parameter <math>k_{on6}</math>, in order to explore the full range of plausible values, the mode of the log-normal distribution is set to <math>0.1 nM ^{-1 } min^{-1}</math> and the confidence interval factor is <math> 3 </math>. Thus the range where 95.45% of the values are found is between <math>0.0333</math> and <math>0.3</math> <math>nM ^{-1 } min^{-1}</math>. |
The probability distributions for the two parameters, adjusted accordingly in order to reflect the above values, are the following: | The probability distributions for the two parameters, adjusted accordingly in order to reflect the above values, are the following: | ||
Revision as of 18:00, 8 December 2015
Two ScbR (R) proteins bind together to form an ScbR homo-dimer (R2).
Contents
Chemical equation
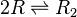
Rate equation
![r= \frac{k^{-}_{6}}{K_{d6}}\cdot [R]^{2} - k^{-}_{6}\cdot [R_{2}]](/wiki/images/math/b/d/2/bd256ae2f0b0944a1e29766d6d3329ee.png)
Parameters
The parameters of this reaction are the dissociation constant for binding of one ScbR to another (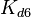 ) and the dissociation rate for binding of one ScbR to another (
) and the dissociation rate for binding of one ScbR to another (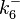 ).
).
| Name | Value | Units | Value in previous GBL models [1] [2] | Remarks-Reference |
|---|---|---|---|---|
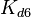
|
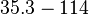 [3] [3]
|
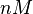
|
N/A | Majka et al. published a study on dimerization of the initiator Protein DnaA of Streptomyces and on its mutants, where they report dissociation constants in the range 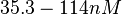 . .
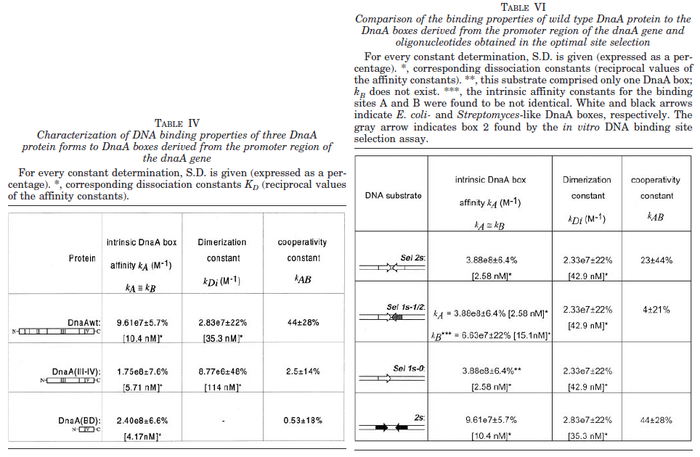 Majka et al. 2001[3] These values agree with Ozbabacan et al. [4] who state that strong protein-protein interactions such as homodimerization have equilibrium dissociation constants |
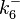
|
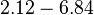 [5] [6] [5] [6]
|

|
N/A | According to Northrup et al. the 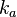 of protein-protein bond formations occur in the order of of protein-protein bond formations occur in the order of 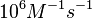 . .
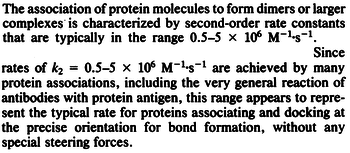 Northrup et al. 1992[6] Therefore, the range Afterwards, the log-normal distribution for the dissociation rate |
Parameters with uncertainty
When deciding how to describe the uncertainty for this parameter we must take into consideration that the values reported in literature correspond to in vitro testing of different protein-protein interaction and dimerization reactions than ScbR, although they refer to another Streptomyces protein (DnaA). This means that there might be a difference between actual parameter values and the ones reported in literature. These facts influence the quantification of the parameter uncertainty and therefore the shape of the corresponding distributions.
More specifically, the weight of the sampling is kept at 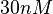 (value that corresponds to the wild type protein homodimerization) which is set as the mode of the log-normal distribution for the
(value that corresponds to the wild type protein homodimerization) which is set as the mode of the log-normal distribution for the 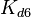 . However, we wish to explore the full nanomolar scale when sampling for parameter values and therefore the confidence interval factor is set to
. However, we wish to explore the full nanomolar scale when sampling for parameter values and therefore the confidence interval factor is set to 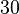 . In this way, the range where 95.45% of the values are found is between
. In this way, the range where 95.45% of the values are found is between 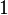 and
and 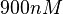 .
.
With regards to the parameter 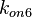 , in order to explore the full range of plausible values, the mode of the log-normal distribution is set to
, in order to explore the full range of plausible values, the mode of the log-normal distribution is set to 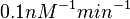 and the confidence interval factor is
and the confidence interval factor is 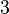 . Thus the range where 95.45% of the values are found is between
. Thus the range where 95.45% of the values are found is between 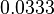 and
and 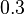
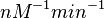 .
.
The probability distributions for the two parameters, adjusted accordingly in order to reflect the above values, are the following:
The location and scale parameters of the distributions are:
| Parameter | μ | σ |
|---|---|---|
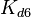
|
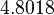
|
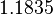
|
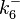
|
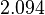
|
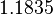
|
References
- ↑ S. Mehra, S. Charaniya, E. Takano, and W.-S. Hu. A bistable gene switch for antibiotic biosynthesis: The butyrolactone regulon in streptomyces coelicolor. PLoS ONE, 3(7), 2008.
- ↑ A. Chatterjee, L. Drews, S. Mehra, E. Takano, Y.N. Kaznessis, and W.-S. Hu. Convergent transcription in the butyrolactone regulon in streptomyces coelicolor confers a bistable genetic switch for antibiotic biosynthesis. PLoS ONE, 6(7), 2011.
- ↑ 3.0 3.1 Majka J, Zakrzewska-Czerwiñska J, Messer W. Sequence recognition, cooperative interaction, and dimerization of the initiator protein DnaA of Streptomyces. J Biol Chem. 2001;276(9):6243-52.
- ↑ Saliha Ece Acuner Ozbabacan, Hatice Billur Engin, Attila Gursoy, and Ozlem Keskin. Transient protein–protein interactions. Protein Engineering, Design and Selection first published online June 15, 2011
- ↑ Janin, Joel. The kinetics of protein-protein recognition. Proteins-Structure Function and Bioinformatics (1997): 153-161.
- ↑ 6.0 6.1 Northrup S.H. and Erickson H.P. Kinetics of protein-protein association explained by Brownian dynamics computer simulation.PNAS 1992;89(8),3338-3342


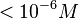 and mostly in the nanomolar range.
and mostly in the nanomolar range.
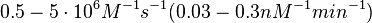 is used to generate the probability distribution of
is used to generate the probability distribution of 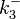 of the ScbR homo-dimer formation is derived from the distributions of
of the ScbR homo-dimer formation is derived from the distributions of