Difference between revisions of "Cellular growth"
(→Dynamics of bacterial colony growth) |
(→Parameters) |
||
| Line 9: | Line 9: | ||
== Parameters == | == Parameters == | ||
| − | The growth rate of the cells is represented by the parameter <math>μ</math>. The parameter values were derived from published data on ''S. coelicolor'' growth rate under different environmental conditions | + | The growth rate of the cells is represented by the parameter <math>μ</math>. In our model, the number of cells is described by a six-parameter Baranyi–Roberts model <ref name="Baranyi1994"> [https://ac.els-cdn.com/0168160594901570/1-s2.0-0168160594901570-main.pdf?_tid=cb6e7472-265f-42d9-aac5-2156512dec23&acdnat=1547635152_30c5eded7fce98579c5a99786f44fd9d Baranyi, J. and T. A. Roberts ''A dynamic approach to predicting bacterial growth in food.'' International Journal of Food Microbiology 23(3): 277-294, 1994.] </ref> <ref name="Baranyi1993"> [https://ac.els-cdn.com/S0740002083710051/1-s2.0-S0740002083710051-main.pdf?_tid=f485f2ad-1765-4c17-839f-895c3f40f88c&acdnat=1547635252_4a7c5bdd80bcca9c47727fc425a281b8 J. Baranyi, T. A. Roberts, P. McClure ''A non-autonomous differential equation to model bacterial growth.'' Food Microbiology 10(1): 43-59, 1993.] </ref> |
| + | The parameter values were derived from published data on ''S. coelicolor'' growth rate under different environmental conditions. | ||
{|class="wikitable" | {|class="wikitable" | ||
Revision as of 11:40, 16 January 2019
Our simulations include hours of cell culture growth in some cases (simulated experimental time, not computational time). Thus, in addition to the dynamics of the regulatory network, we also need to take into account the effects of cell growth. The volume of a single cell is increasing from the beginning of the cell cycle until the moment it divides into two daughter cells and the molecules included in the initial cell are distributed between the two new ones. As a consequence, the cellular growth introduces a dilution in the concentration of the cellular species, which is represented in our deterministic model by adding a dilution term 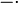 in the
Ordinary Differential Equations (ODEs) of all species, with the exception of the autoinducer in the environment
in the
Ordinary Differential Equations (ODEs) of all species, with the exception of the autoinducer in the environment 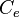 .
.
Parameters
The growth rate of the cells is represented by the parameter Failed to parse (PNG conversion failed; check for correct installation of latex and dvipng (or dvips + gs + convert)): μ . In our model, the number of cells is described by a six-parameter Baranyi–Roberts model [1] [2] The parameter values were derived from published data on S. coelicolor growth rate under different environmental conditions.
| Name | Value | Units | Value in previous GBL models [3] [4] | Remarks-Reference |
|---|---|---|---|---|
| Failed to parse (PNG conversion failed; check for correct installation of latex and dvipng (or dvips + gs + convert)): τ | 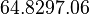 [5][6] [7] [5][6] [7]
|
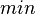
|
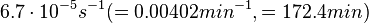 [3] [3]
Range tested: ( Bistability range: ( and ( |
In a study on comparison between turbimetric and gravimetric techniques for the measurements of specific growth rates of streptomycetes, Flowers et al. reported growth rate values in the range of 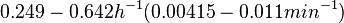 . The reported doubling times were in the range of . The reported doubling times were in the range of 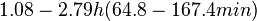 . .
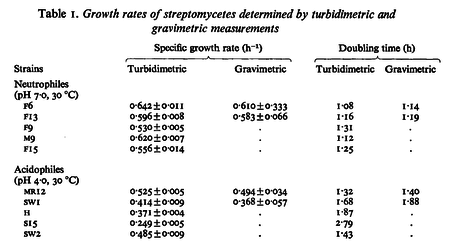 Flowers et al. 1977[5] Additionally, Melzoch et al. studied the observed growth rates of S. coelicolor in variously limited chemostat cultures and reported a maximum growth rate of 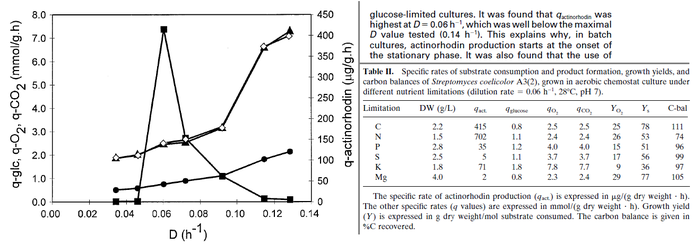 Melzoch et al. 1997[7] These values are comparable with the findings of R.A. Cox, who reported genomic properties and macromolecular compositions of Streptomyces coelicolor A3(2). The largest growth rate was 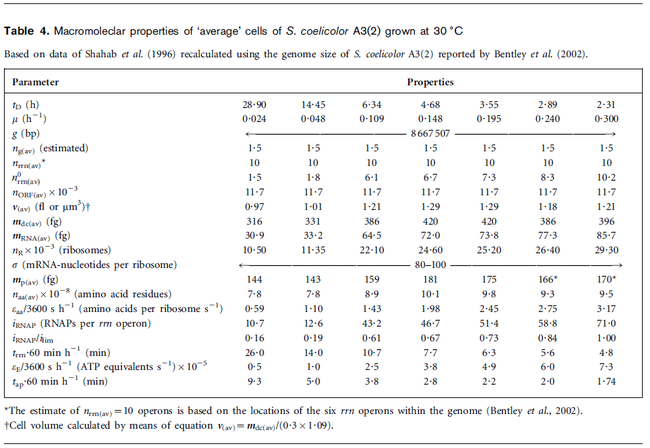 Cox et al. 2004[6] |
Parameters with uncertainty
When deciding how to describe the uncertainty for this parameter we must take into consideration that some of the parameters were calculated or estimated based on experimental data on S. coelicolor cultures, so there might be some error included in the estimations. Additionally, since the simulations will investigate the switch which regulates antibiotic production in S. coelicolor, it would be preferable to be as close as possible to the growth rate which facilitates this process. These facts influence the quantification of the parameter uncertainty and therefore the shape of the corresponding distribution. By assigning the appropriate weights to the parameter values and using the method described here, the appropriate probability distribution was designed.
In order to keep the parameter values as strictly as possible within the range of the values reported in literature, the prediction interval in this case for the definition of the distribution is set to 95.45% rather than 68.27%. Thus, the weight of the distribution will be put to Failed to parse (Cannot store math image on filesystem.): 99.3 min which is also set as the mode of the log-normal distribution for the Failed to parse (PNG conversion failed; check for correct installation of latex and dvipng (or dvips + gs + convert)): μ and the Spread is set to Failed to parse (Cannot store math image on filesystem.): 1.74 , in order to test the system's behaviour under the full range of reported values. This means that the range where 95.45% of the parameter values are found is between Failed to parse (Cannot store math image on filesystem.): 57 and Failed to parse (Cannot store math image on filesystem.): 172.8 min .
The probability distribution for the parameter, adjusted accordingly in order to reflect the above values, is the following:
The values retrieved from literature and their weights are indicated by the blue dashed lines, and the uncertainty for each value is indicated using a default value of 10% error (orange lines).
The parameter information of the distribution is:
| Parameter | Mode | Spread | μ | σ |
|---|---|---|---|---|
| Failed to parse (PNG conversion failed; check for correct installation of latex and dvipng (or dvips + gs + convert)): τ | Failed to parse (Cannot store math image on filesystem.): 99.3 | 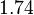
|
Failed to parse (Cannot store math image on filesystem.): 4.6699 | Failed to parse (Cannot store math image on filesystem.): 0.26762 |
References
- ↑ Baranyi, J. and T. A. Roberts A dynamic approach to predicting bacterial growth in food. International Journal of Food Microbiology 23(3): 277-294, 1994.
- ↑ J. Baranyi, T. A. Roberts, P. McClure A non-autonomous differential equation to model bacterial growth. Food Microbiology 10(1): 43-59, 1993.
- ↑ 3.0 3.1 3.2 S. Mehra, S. Charaniya, E. Takano, and W.-S. Hu. A bistable gene switch for antibiotic biosynthesis: The butyrolactone regulon in streptomyces coelicolor. PLoS ONE, 3(7), 2008.
- ↑ 4.0 4.1 4.2 A. Chatterjee, L. Drews, S. Mehra, E. Takano, Y.N. Kaznessis, and W.-S. Hu. Convergent transcription in the butyrolactone regulon in streptomyces coelicolor confers a bistable genetic switch for antibiotic biosynthesis. PLoS ONE, 6(7), 2011.
- ↑ 5.0 5.1 Flowers TH, Williams ST. Measurement of growth rates of streptomycetes: comparison of turbidimetric and gravimetric techniques. J Gen Microbiol. 1977;98(1):285-9.
- ↑ 6.0 6.1 Cox RA. Quantitative relationships for specific growth rates and macromolecular compositions of Mycobacterium tuberculosis, Streptomyces coelicolor A3(2) and Escherichia coli B/r: an integrative theoretical approach. Microbiology. 2004 May;150(Pt 5):1413-26.
- ↑ 7.0 7.1 Melzoch K., Teixeira de Mattos M.J., Neijssel O.M. Production of actinorhodin by Streptomyces coelicolor A3(2) grown in chemostat culture. Biotechnology and Bioengineering 1997;54(6): p. 577–582


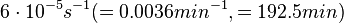
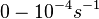
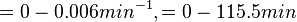 )
)
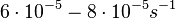
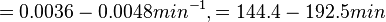 )
)
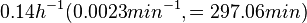
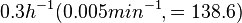 . He noted that the maximum specific growth rate, is attained when cells are amply supplied with the most favourable nutrients.
. He noted that the maximum specific growth rate, is attained when cells are amply supplied with the most favourable nutrients.