Difference between revisions of "46BR.1N fibroblasts Proteomic Analysis Results"
| Line 3: | Line 3: | ||
== Preliminary analysis of the proteomic dataset == | == Preliminary analysis of the proteomic dataset == | ||
| − | The aim of this study was to investigate the proteins related to the AA cascade using SWATH-MS. As this was a preliminary study, the results are presented as raw intensities . | + | The aim of this study was to investigate the proteins related to the AA cascade using SWATH-MS. As this was a preliminary study, the results are presented as raw intensities. |
To investigate the effect of treatment on protein expression, cells stimulated with A23187 (5 µM, 6h), UVR (15 mJ/cm2, 24h) and unstimulated cells were analysed. | To investigate the effect of treatment on protein expression, cells stimulated with A23187 (5 µM, 6h), UVR (15 mJ/cm2, 24h) and unstimulated cells were analysed. | ||
Peptides were detected for all proteins responsible for the production of lipid mediators detected by LC-MS/MS. No peptides were detected for the UVR inducible proteins (COX-2 and mPGES-1) in the irradiated samples. The protein that was detected at the highest mean intensity and the most consistently was PTGR-2, the prostaglandin reductase protein. The most inconsistent protein was PGFS (AKR13), the aldo-keto reductase family 1-member C3 isoform of prostaglandin F synthase. When the different treatments were compared, no difference could be found. | Peptides were detected for all proteins responsible for the production of lipid mediators detected by LC-MS/MS. No peptides were detected for the UVR inducible proteins (COX-2 and mPGES-1) in the irradiated samples. The protein that was detected at the highest mean intensity and the most consistently was PTGR-2, the prostaglandin reductase protein. The most inconsistent protein was PGFS (AKR13), the aldo-keto reductase family 1-member C3 isoform of prostaglandin F synthase. When the different treatments were compared, no difference could be found. | ||
| + | |||
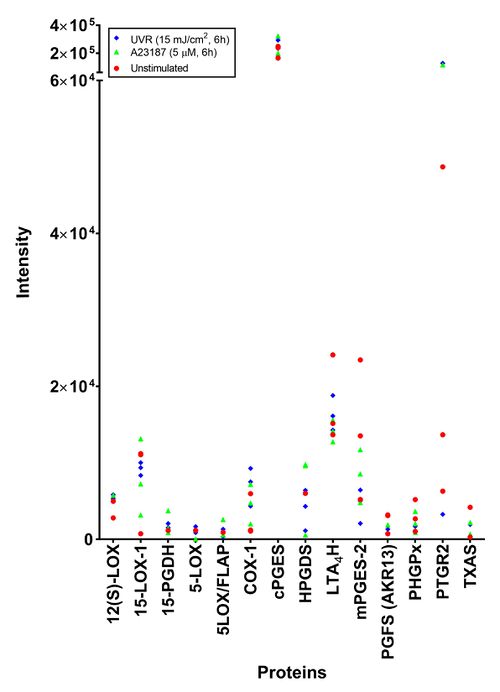
| + | [[Image:F_Proteomics.jpg|none|thumb|500px|Proteomic analysis of AA cascade proteins in 46BR.1N fibroblasts. Results shown as the raw intensities from three independent experiments of unstimulated (red circle), A23187 stimulated (green triangle) and UVR stimulated 46BR.1N fibroblasts (blue diamond).]] | ||
{| class="wikitable" | {| class="wikitable" | ||
| Line 183: | Line 185: | ||
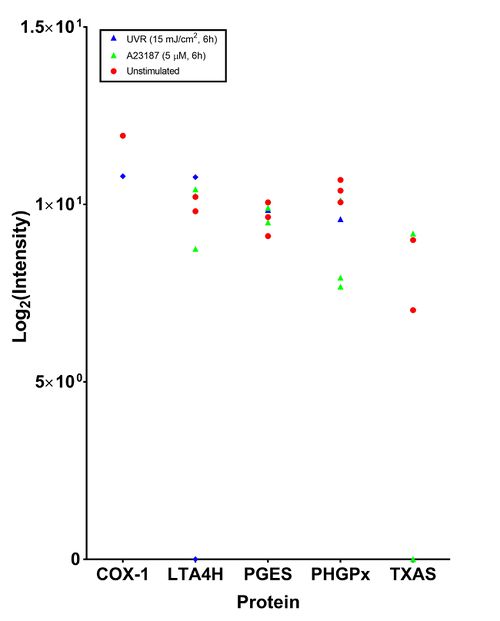
This analysis, performed by the Stoller centre, included taking the top three most intense peptide species, a 1% false discovery rate and acceptance of transition groups if it was identified in the majority of replicates (Table *, Figure *). This in-depth analysis only found five proteins related to the eicosanoid cascade. PHGPx and LTA4H were consistently identified in 7/9 samples. The isoform of PGES which was detected was not specified. | This analysis, performed by the Stoller centre, included taking the top three most intense peptide species, a 1% false discovery rate and acceptance of transition groups if it was identified in the majority of replicates (Table *, Figure *). This in-depth analysis only found five proteins related to the eicosanoid cascade. PHGPx and LTA4H were consistently identified in 7/9 samples. The isoform of PGES which was detected was not specified. | ||
| + | |||
| + | [[Image:Proteomics_F_-_dave.jpg|none|thumb|500px|In-depth proteomic analysis of AA cascade proteins in 46BR.1N fibroblasts. Results shown are from three independent experiments of unstimulated (red circle), A23187 stimulated (green triangle) and UVR stimulated 46BR.1N fibroblasts (blue diamond).]] | ||
| + | |||
| + | {| class="wikitable" | ||
| + | ! rowspan="2" style="font-weight:bold;" | Proteins | ||
| + | ! colspan="3" style="font-weight:bold;" | Unstimulated | ||
| + | ! colspan="3" style="font-weight:bold;" | A23187 (5 µM, 6h) | ||
| + | ! colspan="3" style="font-weight:bold;" | UVR (15 mJ/cm2, 24h) | ||
| + | |- | ||
| + | | style="font-weight:bold;" | Mean | ||
| + | | style="font-weight:bold;" | SD | ||
| + | | style="font-weight:bold;" | n | ||
| + | | style="font-weight:bold;" | Mean | ||
| + | | style="font-weight:bold;" | SD | ||
| + | | style="font-weight:bold;" | n | ||
| + | | style="font-weight:bold;" | Mean | ||
| + | | style="font-weight:bold;" | SD | ||
| + | | style="font-weight:bold;" | n | ||
| + | |- | ||
| + | | COX-1 | ||
| + | | 11.94 | ||
| + | | 0 | ||
| + | | 1 | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | | 10.8 | ||
| + | | 0 | ||
| + | | 1 | ||
| + | |- | ||
| + | | LTA4H | ||
| + | | 10.01 | ||
| + | | 0.29 | ||
| + | | 2 | ||
| + | | 9.81 | ||
| + | | 0.92 | ||
| + | | 3 | ||
| + | | 5.39 | ||
| + | | 7.62 | ||
| + | | 2 | ||
| + | |- | ||
| + | | PGES | ||
| + | | 9.6 | ||
| + | | 0.48 | ||
| + | | 3 | ||
| + | | 9.7 | ||
| + | | 0.29 | ||
| + | | 2 | ||
| + | | 9.85 | ||
| + | | 0 | ||
| + | | 1 | ||
| + | |- | ||
| + | | PHGPx | ||
| + | | 10.38 | ||
| + | | 0.32 | ||
| + | | 3 | ||
| + | | 8.58 | ||
| + | | 1.34 | ||
| + | | 3 | ||
| + | | 9.59 | ||
| + | | 0 | ||
| + | | 1 | ||
| + | |- | ||
| + | | TXAS | ||
| + | | 8.01 | ||
| + | | 1.4 | ||
| + | | 2 | ||
| + | | 3.06 | ||
| + | | 5.3 | ||
| + | | 3 | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |} | ||
Revision as of 07:29, 3 June 2019
Preliminary analysis of the proteomic dataset
The aim of this study was to investigate the proteins related to the AA cascade using SWATH-MS. As this was a preliminary study, the results are presented as raw intensities.
To investigate the effect of treatment on protein expression, cells stimulated with A23187 (5 µM, 6h), UVR (15 mJ/cm2, 24h) and unstimulated cells were analysed. Peptides were detected for all proteins responsible for the production of lipid mediators detected by LC-MS/MS. No peptides were detected for the UVR inducible proteins (COX-2 and mPGES-1) in the irradiated samples. The protein that was detected at the highest mean intensity and the most consistently was PTGR-2, the prostaglandin reductase protein. The most inconsistent protein was PGFS (AKR13), the aldo-keto reductase family 1-member C3 isoform of prostaglandin F synthase. When the different treatments were compared, no difference could be found.
| Unstimulated | A23187 (5 mM, 6h) | UVR (15 mJ/cm2, 6h) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Mean | SD | n | Mean | SD | n | Mean | SD | n | |
| 12(S)-LOX | 3.89E+03 | 1.92E+03 | 4 | 5.77E+03 | 0.00E+00 | 1 | 5.65E+03 | 1.46E+03 | 3 |
| 15-LOX-1 | 8.52E+03 | 5.21E+03 | 4 | 9.18E+03 | 5.18E+03 | 4 | 9.44E+03 | 1.15E+03 | 4 |
| 15-PGDH | 1.16E+03 | 6.06E+02 | 2 | 1.75E+03 | 1.40E+03 | 4 | 1.79E+03 | 3.80E+02 | 2 |
| 5-LOX | 1.17E+03 | 0.00E+00 | 1 | 5.30E+01 | 0.00E+00 | 1 | 1.36E+03 | 1.17E+03 | 4 |
| 5LOX/FLAP | 9.06E+02 | 0.00E+00 | 1 | 1.93E+03 | 1.18E+03 | 3 | 8.32E+02 | 6.92E+02 | 2 |
| COX-1 | 2.36E+03 | 2.40E+03 | 4 | 4.66E+03 | 2.58E+03 | 3 | 6.36E+03 | 2.86E+03 | 4 |
| cPGES | 2.13E+05 | 8.09E+04 | 5 | 2.31E+05 | 1.42E+05 | 6 | 2.30E+05 | 9.48E+04 | 6 |
| HPGDS | 6.01E+03 | 0.00E+00 | 1 | 6.67E+03 | 5.27E+03 | 3 | 4.04E+03 | 2.20E+03 | 4 |
| LTA4H | 1.64E+04 | 5.48E+03 | 5 | 1.42E+04 | 3.14E+03 | 6 | 1.64E+04 | 2.95E+03 | 6 |
| mPGES-2 | 1.22E+04 | 9.04E+03 | 5 | 8.36E+03 | 6.24E+03 | 6 | 4.52E+03 | 2.34E+03 | 6 |
| PGFS (AKR13) | 2.34E+03 | 1.39E+03 | 3 | 1.90E+03 | 0.00E+00 | 1 | 1.32E+03 | 0.00E+00 | 1 |
| PHGPx | 2.90E+03 | 2.18E+03 | 4 | 2.17E+03 | 1.55E+03 | 4 | 1.47E+03 | 4.73E+02 | 3 |
| PTGR2 | 2.29E+04 | 3.68E+04 | 6 | 4.49E+05 | 5.19E+05 | 5 | 2.94E+05 | 5.95E+05 | 6 |
| TXAS | 2.89E+03 | 2.41E+03 | 3 | 1.70E+03 | 1.38E+03 | 3 | 1.94E+03 | 2.02E+03 | 2 |
Further analysis of the proteomic dataset
This analysis, performed by the Stoller centre, included taking the top three most intense peptide species, a 1% false discovery rate and acceptance of transition groups if it was identified in the majority of replicates (Table *, Figure *). This in-depth analysis only found five proteins related to the eicosanoid cascade. PHGPx and LTA4H were consistently identified in 7/9 samples. The isoform of PGES which was detected was not specified.
| Proteins | Unstimulated | A23187 (5 µM, 6h) | UVR (15 mJ/cm2, 24h) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Mean | SD | n | Mean | SD | n | Mean | SD | n | |
| COX-1 | 11.94 | 0 | 1 | 10.8 | 0 | 1 | |||
| LTA4H | 10.01 | 0.29 | 2 | 9.81 | 0.92 | 3 | 5.39 | 7.62 | 2 |
| PGES | 9.6 | 0.48 | 3 | 9.7 | 0.29 | 2 | 9.85 | 0 | 1 |
| PHGPx | 10.38 | 0.32 | 3 | 8.58 | 1.34 | 3 | 9.59 | 0 | 1 |
| TXAS | 8.01 | 1.4 | 2 | 3.06 | 5.3 | 3 | |||