Difference between revisions of "Antisense interaction between r and a"
(→Parameters) |
(→Parameters) |
||
| Line 42: | Line 42: | ||
</ul></div> | </ul></div> | ||
| − | Finally, Nordgren et al. published a study on the kinetics of antisense RNA interactions by using Surface Plasmon Resonance (in wild-type and mutant RNA pairs), where the values reported for the association rate constants are in the range of <math>4.5 \cdot 10^{4}-9.2 \cdot 10^{5} M^{-1} s^{-1}</math> | + | Finally, Nordgren et al. published a study on the kinetics of antisense RNA interactions by using Surface Plasmon Resonance (in wild-type and mutant RNA pairs), where the values reported for the association rate constants are in the range of <math>4.5 \cdot 10^{4}-9.2 \cdot 10^{5} M^{-1} s^{-1}</math> <math>(0.0027-0.0552 nM^{-1} min^{-1})</math> and the resulting equilibrium dissociation constant (calculated from the other parameters and measured by gel shift assay) are in the range of <math>4 \cdot 10^{-10}-8.9 \cdot 10^{-8} M</math> (<math>0.4-89 nM</math>). The experiments measured ''in vitro'' the rate of complex formation and ''in vivo'' the efficiency of control. |
[[Image:Kar-text5.png|none|thumb|700px| Nordgren et al. 2001<ref name="Nordgren2001"></ref>]] | [[Image:Kar-text5.png|none|thumb|700px| Nordgren et al. 2001<ref name="Nordgren2001"></ref>]] | ||
Revision as of 18:08, 19 January 2017
The full length mRNA of scbR (r) binds to the full length mRNA of scbA (a) and form a complex which prevents further translation of both mRNAs.
Contents
Chemical equation
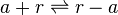
Rate equation
![r= \frac{ k^{-}_{ar}}{K_{ar}}\cdot [r]\cdot [a]-k^{-}_{ar} \cdot [r-a]](/wiki/images/math/0/c/3/0c361c1e3617ab1a9eab319721c9411c.png)
Parameters
The parameters of this reaction is the binding and unbinding rate constant for the interaction of scbR mRNA with scbA mRNA (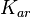 ) and (
) and (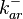 ). The parameter values are derived from various RNA/RNA interaction studies in bacteria.
). The parameter values are derived from various RNA/RNA interaction studies in bacteria.
| Name | Value | Units | Value in previous GBL model [1] | Remarks-Reference |
|---|---|---|---|---|
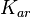
|
Failed to parse (Cannot store math image on filesystem.): 0.40-89 [2] [3] [4] [5] [6] | 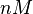
|
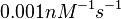 [1] [1]
(Bistability range: |
In a 1991 review, Y. Eguchi reported association rates for double stranded RNA binding in the range of 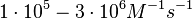 (Failed to parse (Cannot store math image on filesystem.): 0.006-0.18 nM^{-1} min^{-1}
). A similar range (~ (Failed to parse (Cannot store math image on filesystem.): 0.006-0.18 nM^{-1} min^{-1}
). A similar range (~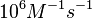 ) is published by S. Brantl in a 2007 review on regulatory mechanisms employed by antisense RNAs. ) is published by S. Brantl in a 2007 review on regulatory mechanisms employed by antisense RNAs.
These values are also in agreement with an in vitro study by Franch et al. on the effect of a U-turn loop structure in RNA/RNA interactions, which employed E. coli hok-RNAs and Sok-RNAs. The study reported binding rate constants Finally, Nordgren et al. published a study on the kinetics of antisense RNA interactions by using Surface Plasmon Resonance (in wild-type and mutant RNA pairs), where the values reported for the association rate constants are in the range of 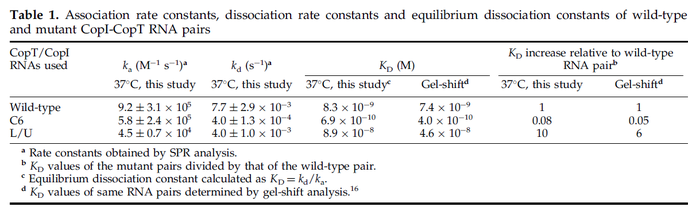 Nordgren et al. 2001[6] Finally, Lima et al. (see table below) in a study measuring kinetics on synthetic oligonucleotides, reported a range of association constants between Therefore, the overall range of values for the association rate |
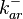
|
Failed to parse (Cannot store math image on filesystem.): 0.006-6 [6] [7] | 
|
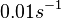 [1] [1]
(Bistability range: |
In the study by Nordgren et al., dissociation rate constants in the range 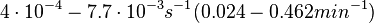 were reported (see figure above). These values are also consistent with the ones published by Lima et al. in a study on antisense oligonucleotide hybridization kinetics, who reported dissociation rate constant values in the range were reported (see figure above). These values are also consistent with the ones published by Lima et al. in a study on antisense oligonucleotide hybridization kinetics, who reported dissociation rate constant values in the range 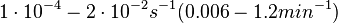
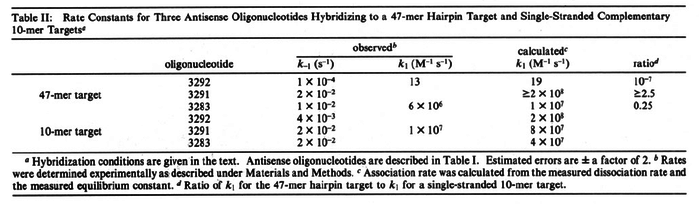 Lima et al. 2001[7] Although the values retrieved by literature are between |
Parameters with uncertainty
When deciding how to describe the uncertainty for these parameter we must take into consideration the fact that many of the reported values are derived from in vitro experiments and correspond to mRNA sequences or fragments of different bacteria species. This means that there might be some difference between actual parameter values and the ones reported in literature. These facts influence the quantification of the parameter uncertainty and therefore the shape of the corresponding distribution.
With regards to the 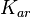 the values that correspond to the wild type RNAs are between
the values that correspond to the wild type RNAs are between 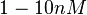 , therefore we put the weight of the distribution in this range and we consider as least likely the larger values as they correspond to mutant RNAs. Therefore, the mode of the log-normal distribution is set to
, therefore we put the weight of the distribution in this range and we consider as least likely the larger values as they correspond to mutant RNAs. Therefore, the mode of the log-normal distribution is set to 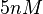 and the confidence interval factor is
and the confidence interval factor is 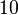 . Thus the range where 95.45% of the values are found is between Failed to parse (Cannot store math image on filesystem.): 0.5
and Failed to parse (Cannot store math image on filesystem.): 50
.
. Thus the range where 95.45% of the values are found is between Failed to parse (Cannot store math image on filesystem.): 0.5
and Failed to parse (Cannot store math image on filesystem.): 50
.
Similarly, the values reported for 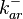 are within the range
are within the range 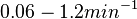 , we put the weight of the distribution within this range but also explore the whole range of values. Therefore, the mode of the log-normal distribution is set to Failed to parse (Cannot store math image on filesystem.): 0.6
and the confidence interval factor is
, we put the weight of the distribution within this range but also explore the whole range of values. Therefore, the mode of the log-normal distribution is set to Failed to parse (Cannot store math image on filesystem.): 0.6
and the confidence interval factor is 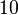 . Thus the range where 95.45% of the values are found is between
. Thus the range where 95.45% of the values are found is between 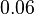 and Failed to parse (Cannot store math image on filesystem.): 6 nM
.
and Failed to parse (Cannot store math image on filesystem.): 6 nM
.
Finally, the probability distribution for 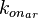 is defined accordingly, in order to allow the exploration of the full range of the values retrieved from literature. Therefore, the mode is set to Failed to parse (Cannot store math image on filesystem.): 0.04 nM^{-1} min^{-1}
and the confidence interval factor is
is defined accordingly, in order to allow the exploration of the full range of the values retrieved from literature. Therefore, the mode is set to Failed to parse (Cannot store math image on filesystem.): 0.04 nM^{-1} min^{-1}
and the confidence interval factor is 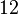 . In this way the range where 95.45% of the values are found is between
. In this way the range where 95.45% of the values are found is between 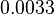 and Failed to parse (Cannot store math image on filesystem.): 0.48 nM^{-1} min^{-1}
. The initial distribution parameters are μ=-2.3296 and σ=0.9431.
and Failed to parse (Cannot store math image on filesystem.): 0.48 nM^{-1} min^{-1}
. The initial distribution parameters are μ=-2.3296 and σ=0.9431.
Since the three parameters are interdependent, thermodynamic consistency also needs to be taken into account. This is achieved by creating a bivariate system as described here. Since 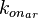 is the parameter with the largest geometric coefficient of variation, this is set as the dependent parameter as per:
is the parameter with the largest geometric coefficient of variation, this is set as the dependent parameter as per: 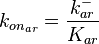 , and an updated probability distribution is defined. The location and scale parameters of
, and an updated probability distribution is defined. The location and scale parameters of 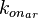 (μ=-2.1202 and σ=1.26021) were calculated from those of
(μ=-2.1202 and σ=1.26021) were calculated from those of 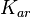 and
and 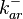 .
.
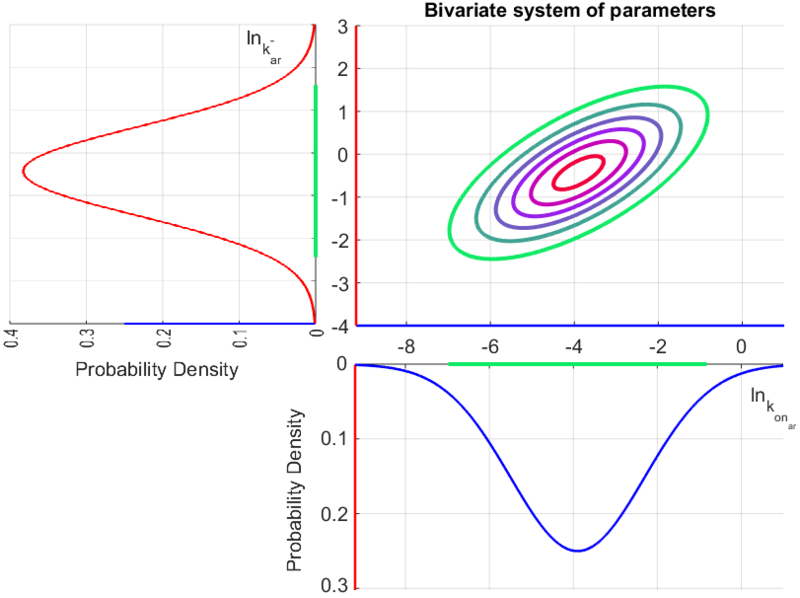
The probability distributions for the three parameters, adjusted accordingly in order to reflect the above values, are the following:
The correlation matrix which is necessary to define the relationship between the two marginal distributions (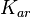 ,
,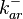 ) of the bivariate system is derived by employing random values generated by the two distributions.
) of the bivariate system is derived by employing random values generated by the two distributions.
The final parameters of the distributions of the multivariate system are:
| Parameter | μ | σ | Correlation matrix |
|---|---|---|---|
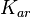
|
Failed to parse (Cannot store math image on filesystem.): 2.4035 | 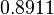
|
N/A |
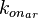
|
Failed to parse (Cannot store math image on filesystem.): -2.1202 | Failed to parse (Cannot store math image on filesystem.): 1.26021 | Failed to parse (Cannot store math image on filesystem.): \begin{pmatrix} 1 & 0.5937 \\ 0.5937 & 1 \end{pmatrix} |
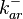
|
Failed to parse (Cannot store math image on filesystem.): 0.2833 | 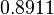
|
The multivariate system of the normal distributions (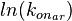 and
and 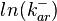 ) and the resulting samples of values are presented in the following figure:
) and the resulting samples of values are presented in the following figure:
In this way, a system of distributions is created where each distribution is described and constrained by the other two. Therefore, the parameters will be sampled by the two marginal distributions in a way consistent with our beliefs and with the relevant thermodynamic constraints. However, since the model's reaction rate requires the parameters 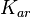 and
and 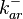 , and not the
, and not the 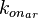 , the value for
, the value for 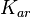 is calculated by the parameters sampled from the other two distributions in an additional step, as per
is calculated by the parameters sampled from the other two distributions in an additional step, as per 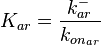 .
.
References
- ↑ 1.0 1.1 1.2 A. Chatterjee, L. Drews, S. Mehra, E. Takano, Y.N. Kaznessis, and W.-S. Hu. Convergent transcription in the butyrolactone regulon in streptomyces coelicolor confers a bistable genetic switch for antibiotic biosynthesis. PLoS ONE, 6(7), 2011.
- ↑ Shokeen S, Johnson CM, Greenfield TJ, Manias DA, Dunny GM, Weaver KE. Structural analysis of the Anti-Q-Qs interaction: RNA-mediated regulation of E. faecalis plasmid pCF10 conjugation. Plasmid. 2010;64(1):26-35.
- ↑ 3.0 3.1 Brantl S. Regulatory mechanisms employed by cis-encoded antisense RNAs. Cell regulation (RNA special issue). 2007; 10: 102–109.
- ↑ 4.0 4.1 Eguchi Y., Itoh T., Tomizawa J. Antisense RNA. Annual Review of Biochemistry. 1991;60: 631–652.
- ↑ 5.0 5.1 5.2 Franch T., Petersen M., Wagner EG., Jacobsen JP., Gerdes K. Antisense RNA regulation in prokaryotes: rapid RNA/RNA interaction facilitated by a general U-turn loop structure. J Mol Biol. 1999 Dec 17;294(5):1115-25.
- ↑ 6.0 6.1 6.2 Nordgren S., Slagter-Jäger J.G., Wagner G.H. Real time kinetic studies of the interaction between folded antisense and target RNAs using surface plasmon resonance. J. Mol. Biol. 2001;310(5):1125-34.
- ↑ 7.0 7.1 Lima W.F., Monia B.P., Ecker D.J., Freier S. M. Implication of RNA structure on antisense oligonucleotide hybridization kinetics. Biochemistry 1992 31 (48), 12055-12061


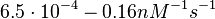 )
)
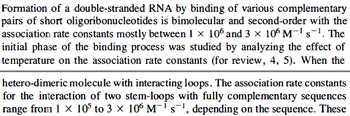
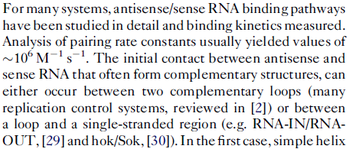


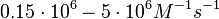 (
(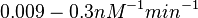 ) and suggested that ~
) and suggested that ~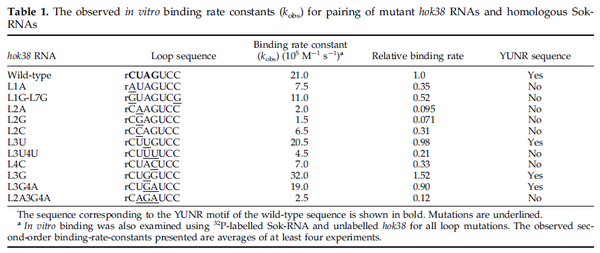
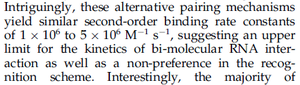

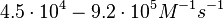
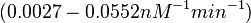 and the resulting equilibrium dissociation constant (calculated from the other parameters and measured by gel shift assay) are in the range of
and the resulting equilibrium dissociation constant (calculated from the other parameters and measured by gel shift assay) are in the range of 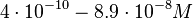 (
(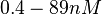 ). The experiments measured in vitro the rate of complex formation and in vivo the efficiency of control.
). The experiments measured in vitro the rate of complex formation and in vivo the efficiency of control.
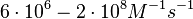 (
(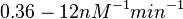 ).
).
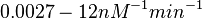 and for the equilibrium dissociation constant
and for the equilibrium dissociation constant 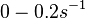 )
)
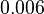 and
and 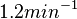 , we wish to explore a wider range of values (
, we wish to explore a wider range of values (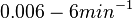 ) in order to take into account the possibility that high GC contents may affect the kinetic rate constants.
) in order to take into account the possibility that high GC contents may affect the kinetic rate constants.