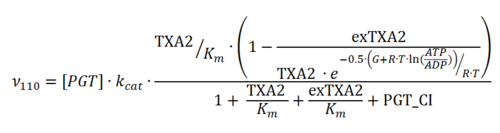Difference between revisions of "Transformation of TXA2 to exTXA2 by PGT"
(→Transport Parameters) |
(→Transport Parameters) |
||
| Line 102: | Line 102: | ||
|1024 | |1024 | ||
|<ref name="SHIGEKAZU1996"> [http://molpharm.aspetjournals.org/content/molpharm/50/4/736.full.pdf Shigekazu I. ''Structural Determinants of Substrates for the Prostaglandin Transporter PGT'' Molecular Pharmacology October 1996, 50 (4) 736-742; DOI: https://doi.org/10.1124/mol.50.4.736]</ref> | |<ref name="SHIGEKAZU1996"> [http://molpharm.aspetjournals.org/content/molpharm/50/4/736.full.pdf Shigekazu I. ''Structural Determinants of Substrates for the Prostaglandin Transporter PGT'' Molecular Pharmacology October 1996, 50 (4) 736-742; DOI: https://doi.org/10.1124/mol.50.4.736]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===k<sub>cat</sub>=== | ||
| + | No data available on the turnover of PGT/ an organic molecule transporter therefore based upon estimates made by http://book.bionumbers.org/what-are-the-rates-of-membrane-transporters/ | ||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Turnover Number | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Transporter | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |3.5 | ||
| + | |min-1 | ||
| + | |Glucose transporter ptsI | ||
| + | |Organism: Bacteria Escherichia coli | ||
| + | Back of the envelope calculation by BioNumbers: http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=102931 | ||
| + | |8 | ||
| + | |<ref name="Waygood1980"> [https://www.ncbi.nlm.nih.gov/pubmed/6992959?dopt=Abstract Waygood EB ''Enzyme I of the phosphoenolpyruvate: sugar phosphotransferase system of Escherichia coli. Purification to homogeneity and some properties.''Can J Biochem. 1980 Jan;58(1):40-8.]</ref> | ||
| + | |- | ||
| + | |0.33 - 0.83 | ||
| + | |min-1 | ||
| + | |Rate of transport of H+/Lactose transporter | ||
| + | |Organism: Bacteria Escherichia coli | ||
| + | Back of the envelop calculation on bionumbers http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=103159 | ||
| + | |8 | ||
| + | |<ref name="Naftalin2007"> [https://www.ncbi.nlm.nih.gov/pubmed/17325012?dopt=Abstract Naftalin R. J. ''ELactose permease H+-lactose symporter: mechanical switch or Brownian ratchet?''Biophys J. 2007 May 15;92(10):3474-91. Epub 2007 Feb 26.]</ref> | ||
| + | |- | ||
| + | |3.28 | ||
| + | |min-1 | ||
| + | |Catalytic Rate of transporter HXT7 | ||
| + | |Organism: Budding yeast Saccharomyces cerevisiae | ||
| + | Back of the envelope calculation http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=101737 | ||
| + | |8 | ||
| + | |<ref name="Ye2001"> [https://www.ncbi.nlm.nih.gov/pubmed/11561293?dopt=Abstract Ye L. ''Expression and activity of the Hxt7 high-affinity hexose transporter of Saccharomyces cerevisiae''Yeast. 2001 Sep 30;18(13):1257-67.]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGT kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.33E+00 || 7.30E+00 || 9.20E-01 || 8.00E-01 | ||
| + | |} | ||
| + | |||
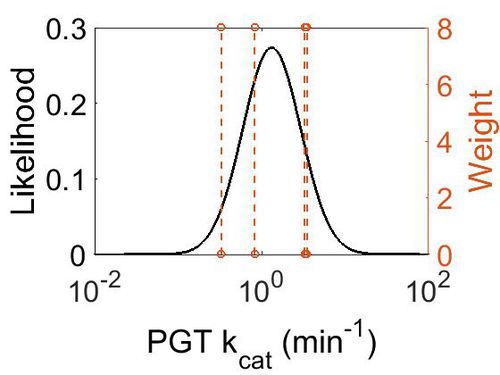
| + | [[Image:132.jpg|none|thumb|500px|The estimated probability distribution for PGT kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === Transporter concentration === | ||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Abundance | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |125 | ||
| + | |ppm | ||
| + | |Human | ||
| + | |Vector:Lung | ||
| + | Enzyme: SLCO2A1 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |2048 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
| + | |- | ||
| + | |17.4 | ||
| + | |ppm | ||
| + | |Human | ||
| + | |Expression Vector: Esophagus | ||
| + | Enzyme: SLCO2A1 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |2048 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |2.82 | ||
| + | |ppm | ||
| + | |Human | ||
| + | |Vector:Gut | ||
| + | Enzyme: SLCO2A1 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |2048 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
|} | |} | ||
Revision as of 16:22, 22 May 2019
Contents
Rate equation
Transport Parameters
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 0.0054 | mM | Human Eye | Vector: HEK-OATP2A1 or HEK-OATP2B1 and the respective vector controls
Substrate: Latanoprost, a prostaglandin F2a analogue pH: 7.5 Temp: 37 °C |
2048 | [1] |
| 0.000331 ± 0.000131 | mM | Transfected Human embryonic kidney cells (HEK293) | Substrate: PGE2
pH: unknown Temp: 37 °C |
512 | [2] |
| 0.007202 ± 0.000595 | mM | Transfected Human embryonic kidney cells (HEK293) | Substrate: PGE3
pH: unknown Temp: 37 °C |
512 | [2] |
| 0.000376 ± 3.4e-5 | mM | PGT-expressing Madin-Darby Canine Kidney Epithelial Cells (MDCK) cells | Substrate: PGH2
pH: unknown Temp: unknown |
192 | [3] |
| 9.40E-05 | mM | Human | Substrate: PGE2
Species: Human Vector: HeLa Cell Line pH: 7 Temperature: 27 |
1024 | [4] |
| 0.000423 | mM | Human | Substrate: TXB2
Species: Human Vector: HeLa Cell Line pH: 7 Temperature: 27 |
1024 | [4] |
| 0.007569 | mM | Human | Substrate: 6-Keto-PGF1a
Species: Human Vector: HeLa Cell Line pH: 7 Temperature: 27 |
1024 | [4] |
| 4.87E-05 | mM | Human | Substrate: PGE2
Species: Human Vector: HeLa Cell Line pH: 7 Temperature: 27 |
1024 | [5] |
kcat
No data available on the turnover of PGT/ an organic molecule transporter therefore based upon estimates made by http://book.bionumbers.org/what-are-the-rates-of-membrane-transporters/
| Value | Units | Transporter | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 3.5 | min-1 | Glucose transporter ptsI | Organism: Bacteria Escherichia coli
Back of the envelope calculation by BioNumbers: http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=102931 |
8 | [6] |
| 0.33 - 0.83 | min-1 | Rate of transport of H+/Lactose transporter | Organism: Bacteria Escherichia coli
Back of the envelop calculation on bionumbers http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=103159 |
8 | [7] |
| 3.28 | min-1 | Catalytic Rate of transporter HXT7 | Organism: Budding yeast Saccharomyces cerevisiae
Back of the envelope calculation http://bionumbers.hms.harvard.edu/bionumber.aspx?&id=101737 |
8 | [8] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.33E+00 | 7.30E+00 | 9.20E-01 | 8.00E-01 |
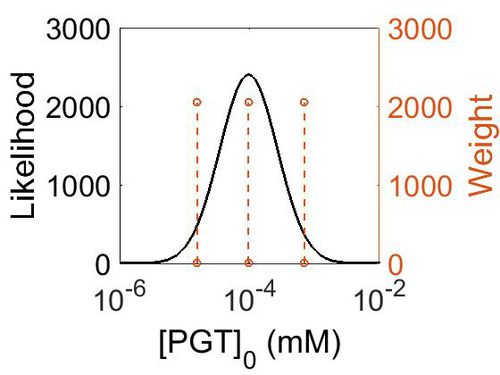
Transporter concentration
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 125 | ppm | Human | Vector:Lung
Enzyme: SLCO2A1 pH: 7.5 Temperature: 37 °C |
2048 | [9] |
| 17.4 | ppm | Human | Expression Vector: Esophagus
Enzyme: SLCO2A1 pH: 7.5 Temperature: 37 °C |
2048 | [10] |
| 2.82 | ppm | Human | Vector:Gut
Enzyme: SLCO2A1 pH: 7.5 Temperature: 37 °C |
2048 | [9] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.73E+01 | 4.72E+00 | 3.90E+00 | 1.02E+00 |
References
- ↑ M. Kraft The prostaglandin transporter OATP2A1 is expressed in human ocular tissues and transports the antiglaucoma prostanoid latanoprost. Invest Ophthalmol Vis Sci. 2010 May;51(5):2504-11. doi: 10.1167/iovs.09-4290. Epub 2009 Dec 17.
- ↑ 2.0 2.1 Gose T. Prostaglandin transporter (OATP2A1/SLCO2A1) contributes to local disposition of eicosapentaenoic acid-derived PGE3. Prostaglandins Other Lipid Mediat. 2016 Jan;122:10-7. doi: 10.1016/j.prostaglandins.2015.12.003. Epub 2015 Dec 10.
- ↑ Chi Y. The prostaglandin transporter PGT transports PGH2 Biochemical and Biophysical Research Communications, Volume 395, Issue 2, 30 April 2010, Pages 168–172
- ↑ 4.0 4.1 4.2 [http://science.sciencemag.org/content/sci/268/5212/866.full.pdf Kanai N. Identification and Characterization of a Prostaglandin Transporter Science 12 May 1995:Vol. 268, Issue 5212, pp. 866-869, DOI: 10.1126/science.7754369]
- ↑ Shigekazu I. Structural Determinants of Substrates for the Prostaglandin Transporter PGT Molecular Pharmacology October 1996, 50 (4) 736-742; DOI: https://doi.org/10.1124/mol.50.4.736
- ↑ Waygood EB Enzyme I of the phosphoenolpyruvate: sugar phosphotransferase system of Escherichia coli. Purification to homogeneity and some properties.Can J Biochem. 1980 Jan;58(1):40-8.
- ↑ Naftalin R. J. ELactose permease H+-lactose symporter: mechanical switch or Brownian ratchet?Biophys J. 2007 May 15;92(10):3474-91. Epub 2007 Feb 26.
- ↑ Ye L. Expression and activity of the Hxt7 high-affinity hexose transporter of Saccharomyces cerevisiaeYeast. 2001 Sep 30;18(13):1257-67.
- ↑ 9.0 9.1 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587