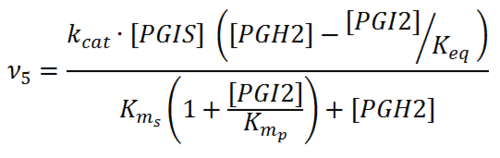Difference between revisions of "Transformation of PGH2 to PGI2"
(→Parameters) |
(→Parameters) |
||
| Line 24: | Line 24: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 33: | Line 34: | ||
pH:7.4 | pH:7.4 | ||
Temperature: 23 | Temperature: 23 | ||
| + | |256 | ||
|<ref name="Yeh2005"> [http://www.ncbi.nlm.nih.gov/pubmed/16406803 H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase.'' Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.]</ref> | |<ref name="Yeh2005"> [http://www.ncbi.nlm.nih.gov/pubmed/16406803 H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase.'' Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.]</ref> | ||
|- | |- | ||
| Line 42: | Line 44: | ||
pH:7.4 | pH:7.4 | ||
Temperature: 24 | Temperature: 24 | ||
| + | |192 | ||
|<ref name="Hara1994"> [http://www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | |<ref name="Hara1994"> [http://www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | ||
|- | |- | ||
| Line 76: | Line 79: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 85: | Line 89: | ||
pH:7.4 | pH:7.4 | ||
Temperature: 24 | Temperature: 24 | ||
| + | |192 | ||
| <ref name="Hara1994"> [www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S. , Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | | <ref name="Hara1994"> [www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S. , Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | ||
|} | |} | ||
| Line 106: | Line 111: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 115: | Line 121: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 124: | Line 131: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 133: | Line 141: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 142: | Line 151: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
|- | |- | ||
| Line 165: | Line 175: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 176: | Line 187: | ||
pH: 7.3 | pH: 7.3 | ||
ionic strength: 0.25 | ionic strength: 0.25 | ||
| + | |64 | ||
|<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=PROSTAGLANDIN-I-SYNTHASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=PROSTAGLANDIN-I-SYNTHASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|- | |- | ||
Revision as of 10:32, 22 May 2019
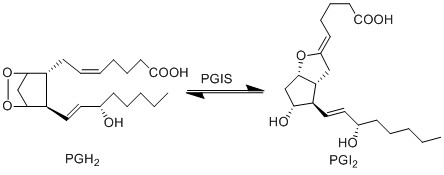
PGH2 is metabolised into the prostacyclin PGI2, by PGIS. This species has been associated with influencing the permeability of vascular compartment (Murata, Ushikubi et al. 1997). Similarly to the TX species, prostacyclins are only found in low concentration and are associated with infiltrating cells from the vascular compartment (Sugimoto, Arai et al. 2006).
Contents
Reaction
Chemical equation
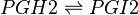
Rate equation
Parameters
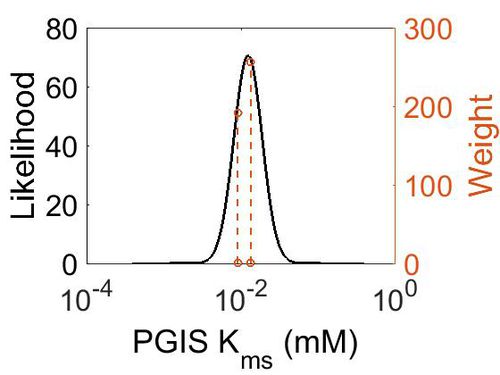
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 1.33E-02 ± 1.40E-03 | 
|
Human | Expression Vector: Bovine Endothelial and Aorta Cells
Enzyme: Human PGIS pH:7.4 Temperature: 23 |
256 | [1] |
| 9.00E-03 ± 5.00E-03 | 
|
Bovine | Expression Vector: Bovine Endothelial and Aorta Cells
Enzyme: Bovine PGIS pH:7.4 Temperature: 24 |
192 | [2] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.24E-02 | 2.46E+00 | -4.22E+00 | 4.20E-01 |
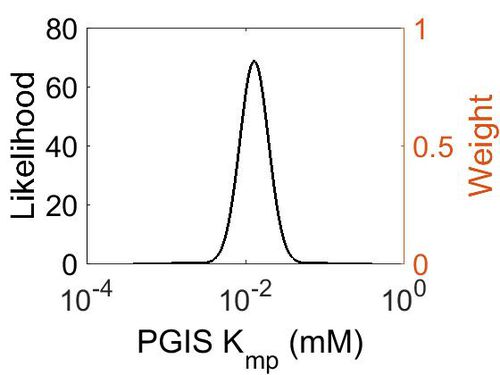
Kmp
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 1.29E-02 | -4.182022605 | 0.414183521 |
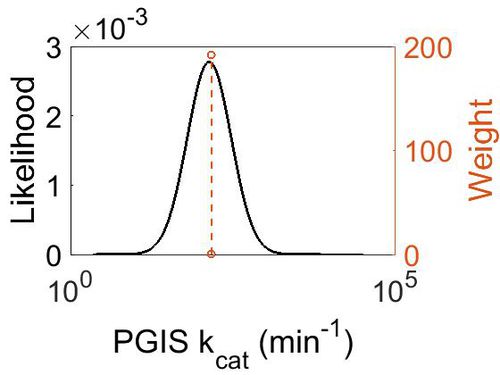
kcat
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 147 ± 45 | per minute | Cattle | Expression Vector: E. Coli
Enzyme: Bovine PGIS pH:7.4 Temperature: 24 |
192 | [2] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.41E+02 | 1.35E+00 | 5.028213628 | 0.287348692 |
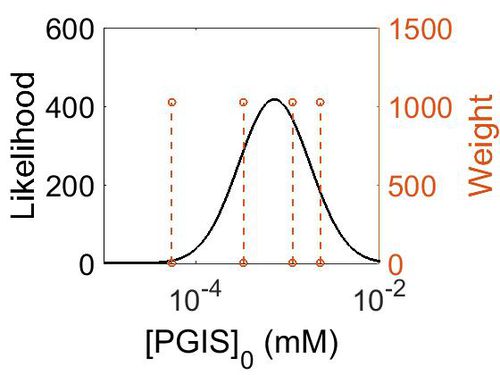
Enzyme concentration
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 412 | 
|
Human | Expression Vector: Urinary bladder
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [3] |
| 206 | 
|
Human | Expression Vector: Lung
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [3] |
| 60.1 | 
|
Human | Expression Vector: Esophagus
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [3] |
| 9.93 | 
|
Human | Expression Vector: Oral Cavity
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.13E+02 | 4.13E+00 | 5.67E+00 | 9.68E-01 |
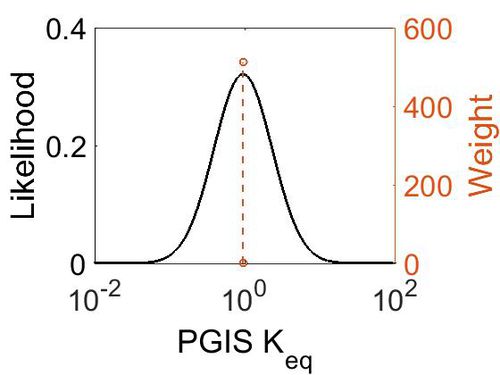
Keq
| Gibbs Free Energy Change | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 0.04 | kcal/mol | Not stated | Estimated
Enzyme: Transacylase Substrate: Product: pH: 7.3 ionic strength: 0.25 |
64 | [5] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 9.35E-01 | 1.00E+01 | 7.30E-01 | 8.90E-01 |
References
- ↑ H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase. Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.
- ↑ 2.0 2.1 Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells. J Biol Chem. 1994 Aug 5;269(31):19897-903. Cite error: Invalid
<ref>tag; name "Hara1994" defined multiple times with different content - ↑ 3.0 3.1 3.2 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471