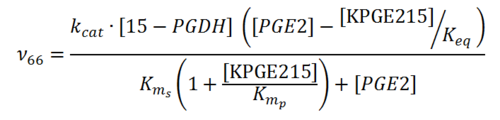Difference between revisions of "Transformation of PGE2 to 15-Keto-PGE2"
(→Enzyme Parameters) |
|||
| Line 25: | Line 25: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 34: | Line 35: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGE2 + NAD+ | Substrate: PGE2 + NAD+ | ||
| + | |1024 | ||
|<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | |<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | ||
|- | |- | ||
| Line 43: | Line 45: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGE2 + NADP+ | Substrate: PGE2 + NADP+ | ||
| + | |1024 | ||
|<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | |<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | ||
|- | |- | ||
| Line 52: | Line 55: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGF2a + NAD+ | Substrate: PGF2a + NAD+ | ||
| + | |1024 | ||
|<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | |<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | ||
|- | |- | ||
| Line 61: | Line 65: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGF2a + NADP+ | Substrate: PGF2a + NADP+ | ||
| + | |1024 | ||
|<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | |<ref name="Fincham1982"> [https://www.ncbi.nlm.nih.gov/pubmed/6575778 N. Fincham, Novel prostaglandin dehydrogenase in rat skin.'' Biochem J. 1983 Apr 15;212(1):129-34.]</ref> | ||
|- | |- | ||
| Line 71: | Line 76: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGE2 | Substrate: PGE2 | ||
| + | |1024 | ||
|<ref name="Zhou2001"> [http://onlinelibrary.wiley.com/doi/10.1046/j.1432-1327.2001.02218.x/epdf Zhou H., C-Terminal region of human NAD+-dependent 15-hydroxyprostaglandin dehydrogenase is involved in the interaction with prostaglandin substrates.'' Eur J Biochem. 2001 Jun;268(12):3368-74.]</ref> | |<ref name="Zhou2001"> [http://onlinelibrary.wiley.com/doi/10.1046/j.1432-1327.2001.02218.x/epdf Zhou H., C-Terminal region of human NAD+-dependent 15-hydroxyprostaglandin dehydrogenase is involved in the interaction with prostaglandin substrates.'' Eur J Biochem. 2001 Jun;268(12):3368-74.]</ref> | ||
|- | |- | ||
| Line 81: | Line 87: | ||
Temperature: 37'C | Temperature: 37'C | ||
Substrate: PGE2 | Substrate: PGE2 | ||
| + | |512 | ||
|<ref name="Zhou2001"> [http://onlinelibrary.wiley.com/doi/10.1046/j.1432-1327.2001.02218.x/epdf Zhou H., C-Terminal region of human NAD+-dependent 15-hydroxyprostaglandin dehydrogenase is involved in the interaction with prostaglandin substrates.'' Eur J Biochem. 2001 Jun;268(12):3368-74.]</ref> | |<ref name="Zhou2001"> [http://onlinelibrary.wiley.com/doi/10.1046/j.1432-1327.2001.02218.x/epdf Zhou H., C-Terminal region of human NAD+-dependent 15-hydroxyprostaglandin dehydrogenase is involved in the interaction with prostaglandin substrates.'' Eur J Biochem. 2001 Jun;268(12):3368-74.]</ref> | ||
|- | |- | ||
| Line 91: | Line 98: | ||
Temperature: 25 | Temperature: 25 | ||
Substrate: PGE2 + NAD+ | Substrate: PGE2 + NAD+ | ||
| + | |1024 | ||
|<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | |<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | ||
|- | |- | ||
| Line 122: | Line 130: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 132: | Line 141: | ||
Temperature: 25 | Temperature: 25 | ||
Substrate: PGE2 + NAD+ | Substrate: PGE2 + NAD+ | ||
| + | |128 | ||
|<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | |<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | ||
|- | |- | ||
| Line 142: | Line 152: | ||
Temperature: 25 | Temperature: 25 | ||
Substrate: PGE2 + NAD+ + Inhibitor | Substrate: PGE2 + NAD+ + Inhibitor | ||
| + | |128 | ||
|<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | |<ref name="Niesen2010"> [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0013719 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships'' PLoS One. 2010 Nov 2;5(11):e13719.]</ref> | ||
|- | |- | ||
| Line 163: | Line 174: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 172: | Line 184: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
|- | |- | ||
| Line 181: | Line 194: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 190: | Line 204: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 199: | Line 214: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |2048 | ||
|Paxdb - Unknown | |Paxdb - Unknown | ||
|} | |} | ||
| Line 219: | Line 235: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 230: | Line 247: | ||
pH: 7.3 | pH: 7.3 | ||
ionic strength: 0.25 | ionic strength: 0.25 | ||
| + | |64 | ||
|<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=ARACHIDONATE-12-LIPOXYGENASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=ARACHIDONATE-12-LIPOXYGENASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
Revision as of 15:57, 22 May 2019
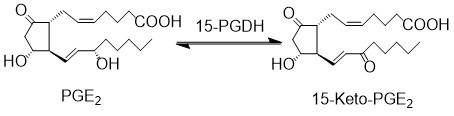
15-hydroxyprostaglandin dehydrogenase, also known as (15-PGDH) metabolises PG into the 15-Keto variant via the oxidation of 15(S)-hydroxyl group. The enzyme is constituently expressed in the skin (Finhelm1982) and within the eicosanoid network it is reported as metabolising PGE2 and PGF2a.
It should be noted that Judson et al found that 15-PGDH expression of the enzyme decreases in response to UVR (Judson2010).
Contents
Reaction
Chemical equation
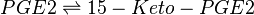
Rate equation
Enzyme Parameters
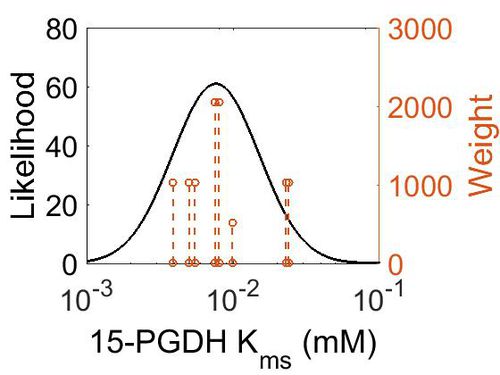
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 0.008 | 
|
Rat Skin | Method:
pH:7.4 Temperature: 37'C Substrate: PGE2 + NAD+ |
1024 | [1] |
| 0.0075 | 
|
Rat Skin | Method:
pH:7.4 Temperature: 37'C Substrate: PGE2 + NADP+ |
1024 | [1] |
| 0.024 | 
|
Rat Skin | Method:
pH:7.4 Temperature: 37'C Substrate: PGF2a + NAD+ |
1024 | [1] |
| 0.023 | 
|
Rat Skin | Method:
pH:7.4 Temperature: 37'C Substrate: PGF2a + NADP+ |
1024 | [1] |
| 0.0039 | 
|
Purified Human 15-PGDH | Method: In vitro
Expression Vector: E. coli pH:7.5 Temperature: 37'C Substrate: PGE2 |
1024 | [2] |
| 0.0099 | 
|
Purified Rat 15-PGDH | Method: In vitro
Expression Vector: E. coli pH:7.5 Temperature: 37'C Substrate: PGE2 |
512 | [2] |
| 0.0055 ± 0.0006 | 
|
Human | Method:In vitro
Expression Vector: E. Coli pH:8 Temperature: 25 Substrate: PGE2 + NAD+ |
1024 | [3] |
| Mode (mM) | Confidence Interval | Location parameter (μ) | Scale parameter (σ) |
|---|---|---|---|
| 7.64E-03 | 5.05E+00 | -4.41E+00 | 6.80E-01 |
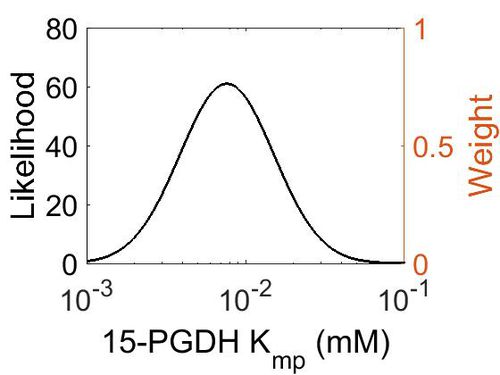
Kmp
| Mode (mM) | Location parameter (μ) | Scale parameter (σ) |
|---|---|---|
| 7.70E-03 | -4.41E+00 | 6.72E-01 |
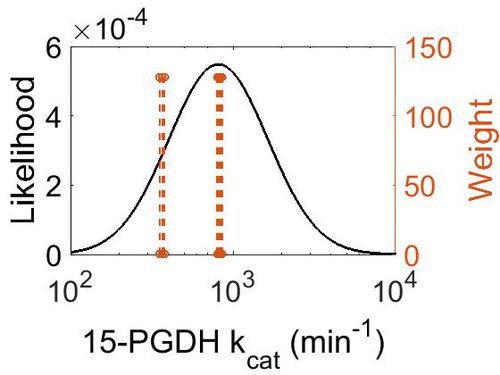
kcat
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 816 ± 18 | 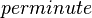
|
Human | Method: In vitro
Expression Vector: E. Coli pH:8 Temperature: 25 Substrate: PGE2 + NAD+ |
128 | [3] |
| 366 ± 12 - 846 ± 12 | 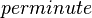
|
Human | Method: In vitro
Expression Vector: E. Coli pH:8 Temperature: 25 Substrate: PGE2 + NAD+ + Inhibitor |
128 | [3] |
| Mode (min-1) | Confidence Interval | Location parameter (μ) | Scale parameter (σ) |
|---|---|---|---|
| 8.12E+02 | 5.38E+00 | 7.19E+00 | 7.01E-01 |
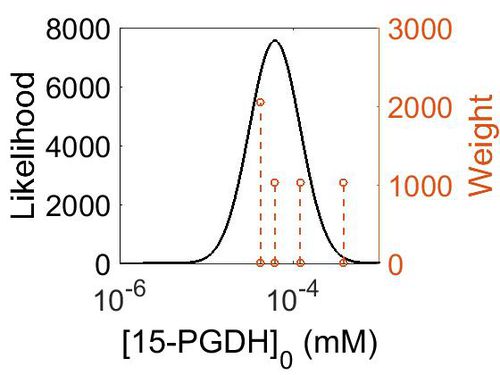
Enzyme concentration
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 69.8 | 
|
Human | Expression Vector: Esophagus
Enzyme: 15-PGDH pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| 22.1 | 
|
Human | Expression Vector: Heart
Enzyme: 15-PGDH pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| 11.2 | 
|
Human | Expression Vector: Esophagus
Enzyme: 15-PGDH pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| 7.68 | 
|
Human | Expression Vector: Skin
Enzyme: 15-PGDH pH: 7.5 Temperature: 37 °C |
2048 | Paxdb - Unknown |
| Mode (mM) | Confidence Interval | Location parameter (μ) | Scale parameter (σ) |
|---|---|---|---|
| 1.12E+01 | 2.32E+00 | 2.87E+00 | 6.80E-01 |
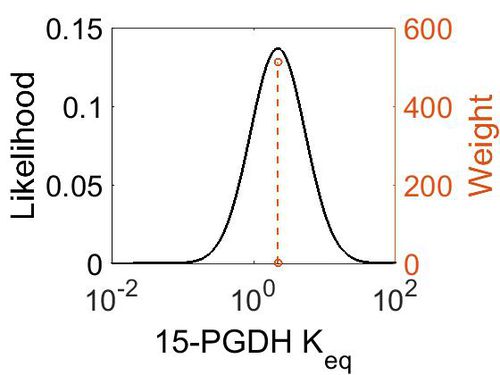
Keq
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| (-0.46818542) | kcal/mol | Calculated | Estimated
Enzyme: 15-PGDH Substrate: PGE2 Product: 15-dehydro-PGE2 pH: 7.3 ionic strength: 0.25 |
64 | [6] |
| Mode | Confidence Interval | Location parameter (μ) | Scale parameter (σ) |
|---|---|---|---|
| 2.21E+00 | 1.00E+01 | 1.58E+00 | 8.91E-01 |
References
- ↑ 1.0 1.1 1.2 1.3 N. Fincham, Novel prostaglandin dehydrogenase in rat skin. Biochem J. 1983 Apr 15;212(1):129-34.
- ↑ 2.0 2.1 Zhou H., C-Terminal region of human NAD+-dependent 15-hydroxyprostaglandin dehydrogenase is involved in the interaction with prostaglandin substrates. Eur J Biochem. 2001 Jun;268(12):3368-74.
- ↑ 3.0 3.1 3.2 F. Niesen,, High-Affinity Inhibitors of Human NAD+-Dependent 15-Hydroxyprostaglandin Dehydrogenase: Mechanisms of Inhibition and Structure-Activity Relationships PLoS One. 2010 Nov 2;5(11):e13719.
- ↑ M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ 5.0 5.1 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471