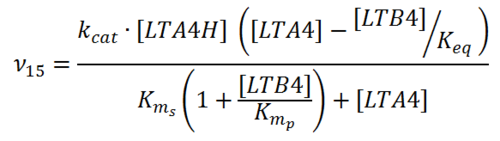Difference between revisions of "Transformation of LTA4 to LTB4"
| Line 14: | Line 14: | ||
== Parameters == | == Parameters == | ||
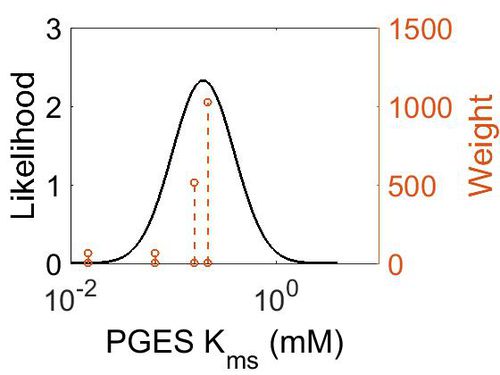
| + | === K<sub>ms</sub> === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 58: | Line 59: | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the LTA4H Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 2.40E-03 || 5.77E+00 || -5.51E+00 || 7.20E-01 | ||
| + | |} | ||
| + | |||
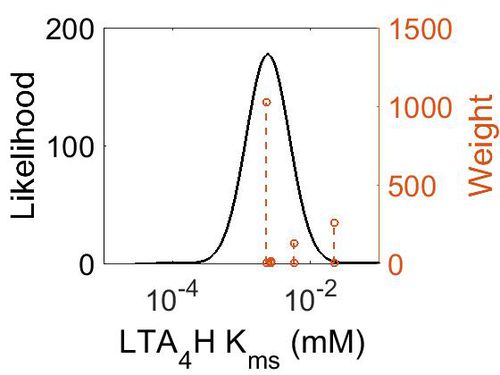
| + | [[Image:49.jpg|none|thumb|500px|The estimated probability distribution for LTA4H Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===K<sub>mp</sub>=== | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the LTA4H Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 2.40E-03 || -5.51E+00 || 7.17E-01 | ||
| + | |} | ||
| + | |||
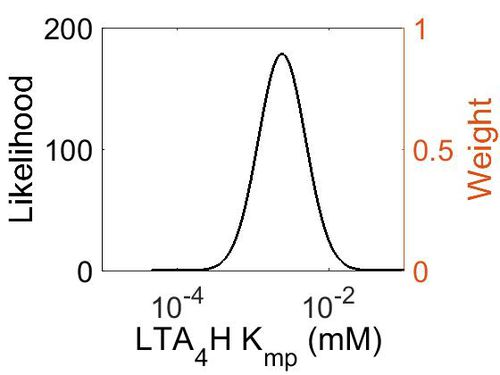
| + | [[Image:50.jpg|none|thumb|500px|The estimated probability distribution for LTA4H Kmp. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
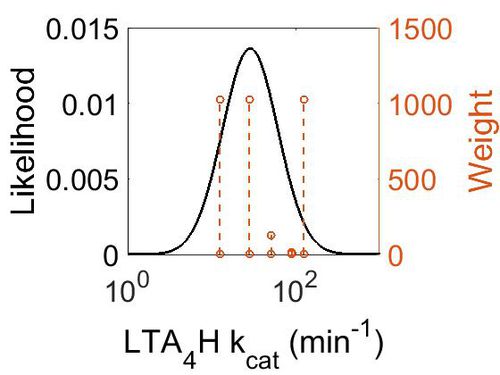
| + | ===k<sub>cat</sub>=== | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 104: | Line 128: | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the LTA4H kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 2.85E+01 || 6.56E+00 || 3.94E+00 || 7.60E-01 | ||
| + | |} | ||
| + | |||
| + | [[Image:51.jpg|none|thumb|500px|The estimated probability distribution for PGES Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===Enzyme concentration=== | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 143: | Line 179: | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the LTA4H concentration distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 4.88E+02 || 1.25E+00 || 6.24E+00 || 2.19E-01 | ||
| + | |} | ||
| + | |||
| + | [[Image:29.jpg|none|thumb|500px|The estimated probability distribution for PGES Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
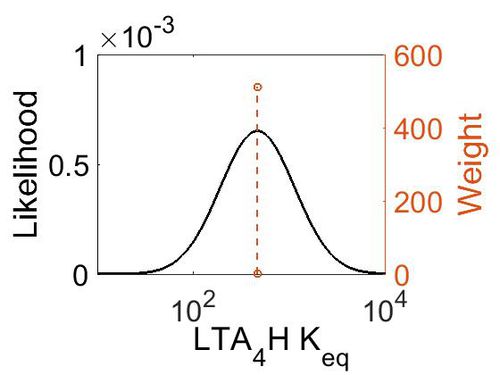
| + | ===K<sub>eq</sub>=== | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
| − | ! | + | ! Gibbs Free Energy Change |
! Units | ! Units | ||
! Species | ! Species | ||
| Line 163: | Line 210: | ||
|<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=LEUKOTRIENE-A4-HYDROLASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=LEUKOTRIENE-A4-HYDROLASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the LTA4H Keq distribution | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 4.64E+02 || 1.00E+01 || 6.93E+00 || 8.90E-01 | ||
| + | |} | ||
| + | |||
| + | [[Image:52.jpg|none|thumb|500px|The estimated probability distribution for PGES Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
== References == | == References == | ||
Revision as of 13:40, 15 May 2019
Contents
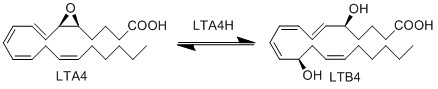
Reaction
Chemical equation
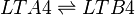
Rate equation
Parameters
Kms
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 5.80E-03 | mM | Human | Expression Vector: E. Coli
Enzyme: Wild Type Leukotriene A4 hydrolase: pH:8 Temperature: 20 (Room Temperature) |
[1] |
| 2.70E-03 | mM | Frog | Expression Vector: oocytes
Enzyme: leukotriene A4 hydrolase pH:8 Temperature: 20 (Room Temperature) |
[2] |
| 2.20E-02 | mM | Human | Expression Vector: Leukocytes
Enzyme: Leukotriene A4 Hydrolase pH: 8 Temperature:2°C |
[3] |
| 2.30E-03 | Expression Vector: Leukocytes
Enzyme: Leukotriene A4 Hydrolase pH: 8 Temperature:37°C |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 2.40E-03 | 5.77E+00 | -5.51E+00 | 7.20E-01 |
Kmp
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 2.40E-03 | -5.51E+00 | 7.17E-01 |
kcat
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 12.6 - 28.2 | per minute | Human | Expression Vector: E. Coli.
Enzyme: Leukotriene A4 Hydrolase pH: 7.4 Temperature: 37 °C |
[4] |
| 51 | per minute | Human | Expression Vector: E. Coli
Enzyme: Wild Type Leukotriene A4 hydrolase: pH:8 Temperature: 20 (Room Temperature) |
[1] |
| 90 | per minute | Frog | Expression Vector: oocytes
Enzyme: leukotriene A4 hydrolase pH:8 Temperature: 20 (Room Temperature) |
[2] |
| 125 | per minute | Human | Expression Vector: Leukocytes
Enzyme: Leukotriene A4 Hydrolase pH: 8 Temperature:37°C |
[3] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 2.85E+01 | 6.56E+00 | 3.94E+00 | 7.60E-01 |
Enzyme concentration
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 714 | 
|
Human | Expression Vector: Lung
Enzyme: LTA4H pH: 7.5 Temperature: 37 °C |
[5] |
| 489 | 
|
Human | Expression Vector: Skin
Enzyme: LTA4H pH: 7.5 Temperature: 37 °C |
[6] |
| 410 | 
|
Human | Expression Vector: Oral Cavity
Enzyme: LTA4H pH: 7.5 Temperature: 37 °C |
[6] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 4.88E+02 | 1.25E+00 | 6.24E+00 | 2.19E-01 |
Keq
| Gibbs Free Energy Change | Units | Species | Notes | Reference |
|---|---|---|---|---|
| (-3.6329994) | kcal/mol | Not stated | Estimated
Enzyme: LTA4H Substrate: LTA4 Product: LTB4 pH: 7.3 ionic strength: 0.25 |
[7] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 4.64E+02 | 1.00E+01 | 6.93E+00 | 8.90E-01 |
References
- ↑ 1.0 1.1 [www.ncbi.nlm.nih.gov/pubmed/8650196 Mueller M. , "Leukotriene A4 hydrolase: protection from mechanism-based inactivation by mutation of tyrosine-378. Proc Natl Acad Sci U S A. 1996 Jun 11;93(12):5931-5.]
- ↑ 2.0 2.1 [www.ncbi.nlm.nih.gov/pubmed/9744798 Stromberg F. , "Purification and characterization of leukotriene A4 hydrolase from Xenopus laevis oocytes. FEBS Lett. 1998 Aug 21;433(3):219-22.]
- ↑ 3.0 3.1 [www.ncbi.nlm.nih.gov/pubmed/6490615 Radmark O. , "Leukotriene A4 hydrolase in human leukocytes. Purification and properties J Biol Chem. 1984 Oct 25;259(20):12339-45.]
- ↑ [www.ncbi.nlm.nih.gov/pubmed/11675384 Rudberg P. , "Leukotriene A4 Hydrolase/Aminopeptidase The Journal of Biological Chemistry, 277, 1398-1404.]
- ↑ M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ 6.0 6.1 M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471