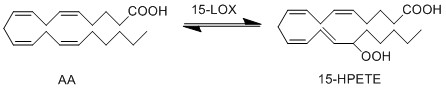
Difference between revisions of "Transformation of AA to 15-HPETE"
(→15-LOX Parameters) |
(→15-LOX Parameters) |
||
| Line 141: | Line 141: | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| + | |} | ||
| + | |||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Gibbs Free Energy Change | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Reference | ||
| + | |- | ||
| + | |(-69.979996) | ||
| + | |kcal/mol | ||
| + | |Not stated | ||
| + | |Estimated | ||
| + | Enzyme: 15-LOX | ||
| + | Substrate: Arachidonate | ||
| + | Product: 15-HPETE | ||
| + | pH: 7.3 | ||
| + | ionic strength: 0.25 | ||
| + | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=RXN66-490 Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
Revision as of 15:36, 18 August 2016
Contents
Reaction
Chemical equation
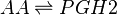
Rate equation
15-LOX Parameters
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 3.70E-03 ± 3.00E-04 | 
|
Human | Expression Vector: Epithelium
Enzyme: 15-Lipoxygenase pH: 7.5 Temperature: 25 |
[1] |
| 5.00E-03 | 
|
Human | Expression Vector: Reticulocyte
Enzyme: 15-Lipoxygenase pH:7.5 Temperature: Unspecified |
[2] |
| 1.06E-02 | 
|
Human | Expression Vector: Keratinocyte
Enzyme: 15-Lipoxygenase pH: 6.7 - 7.3 Temperature: 37 |
[3] |
| 1.17E-02 ± 9.00E-04 | 
|
Human | Expression Vector: Baculovirus Insect Cell
Enzyme: Wildtype 15-Lipoxygenase pH: 6.8 Temperature: 37 |
[4] |
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 595.8 ± 16.8 | 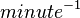
|
Human | Expression Vector: Epithelium
Enzyme: 15-Lipoxygenase pH: 7.5 Temperature: 25 |
[1] |
| 37.2 ± 1.8 | per minute | Expression Vector:Human prostate epithelial
Enzyme:15-lipoxygenase-2 |
pH 7, Temperature: 22 | [5] |
| 45 ± 1.2 | pH 7.5, Temperature: 22 | |||
| 44.4 ± 2.4 | pH 8, Temperature: 22 | |||
| 34.2 ± 0.6 | 15°C, pH = 7.5 | |||
| 45 ± 1.2 | 22°C, pH = 7.5 | |||
| 62.4 ± 4.2 | 30°C, pH = 7.5 | |||
| 82.8 ± 4.8 | 37°C, pH = 7.5 |
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 4.09 | 
|
Human | Expression Vector: Lung
Enzyme: 15-LOX pH: 7.5 Temperature: 37 °C |
[6] |
| 37.4 | 
|
Human | Expression Vector: Spleen
Enzyme: 15-LOX pH: 7.5 Temperature: 37 °C |
[7] |
| 1.40 | 
|
Human | Expression Vector: Gut
Enzyme: 12-LOX pH: 7.5 Temperature: 37 °C |
[6] |
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| (-69.979996) | kcal/mol | Not stated | Estimated
Enzyme: 15-LOX Substrate: Arachidonate Product: 15-HPETE pH: 7.3 ionic strength: 0.25 |
[8] |
References
- ↑ 1.0 1.1 Jacquot C. "Isotope sensitive branching and kinetic isotope effects in the reaction of deuterated arachidonic acids with human 12- and 15-lipoxygenases. Biochemistry. 2008 Jul 8;47(27):7295-303. doi: 10.1021/bi800308q. Epub 2008 Jun 12.
- ↑ Jacquot C. "Synthesis of 11-Thialinoleic Acid and 14-Thialinoleic Acid, Inhibitors of Soybean and Human Lipoxygenases Org Biomol Chem. 2008 Nov 21; 6(22): 4242–4252.
- ↑ Burrall B. "Enzymatic properties of the 15-lipoxygenase of human cultured keratinocytes. J Invest Dermatol. 1988 Oct;91(4):294-7.
- ↑ Sloane D. L. "Conversion of human 15-lipoxygenase to an efficient 12-lipoxygenase: the side-chain geometry of amino acids 417 and 418 determine positional specificity. Protein Eng. 1995 Mar;8(3):275-82..
- ↑ Wecksler A. "Kinetic and Structural Investigations of the Allosteric Site in Human Epithelial 15-Lipoxygenase-2 Biochemistry, 2009, 48 (36), pp 8721–8730
- ↑ 6.0 6.1 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471