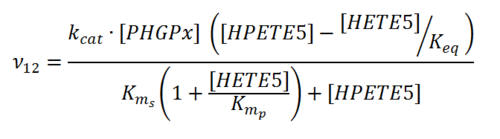Difference between revisions of "Transformation of 5-HPETE to 5-HETE"
| Line 15: | Line 15: | ||
[[File:R12.PNG|center|500px]] | [[File:R12.PNG|center|500px]] | ||
| − | == | + | == Enzyme Parameters == |
| − | + | === K<sub>ms</sub> === | |
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 56: | Line 56: | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PHGPx Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 3.19E-01 || 1.02E+03 || 2.54E+00 || 1.92E+00 | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | === K<sub>mp</sub> === | ||
| + | This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail[[Quantification of parameter uncertainty | here]]). As a result, no confidence interval factor or literature values were cited for this parameter. | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PHGPx Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 3.15E-01 || 2.53E+00 || 1.92E+00 | ||
| + | |} | ||
| + | |||
| + | === k<sub>cat</sub> === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 84: | Line 104: | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PHGPx kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 2.44E+04 || 4.20E+00 || 1.11E+01 || 9.75E-01 | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | === Enzyme concentration === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 131: | Line 162: | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PHGPx concentration distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 3.14E+02 || 1.38E+00 || 5.85E+00 || 3.09E-01 | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | === K<sub>eq</sub> === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
| − | ! | + | ! Gibbs Free Energy Change |
! Units | ! Units | ||
! Species | ! Species | ||
| Line 152: | Line 194: | ||
|<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=1.11.1.12-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=1.11.1.12-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PHGPx Keq distribution | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 5.90E+19 || 1.00E+01 || 4.63E+01 || 8.90E-01 | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
== References == | == References == | ||
Revision as of 10:26, 15 May 2019
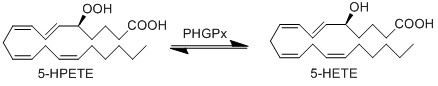
5-HPETE is reduced by phospholipid hydroperoxide glutathione peroxidase (PHGPx) to the corresponding alcohol, 5-HETE (Suth01).
Contents
Reaction
Chemical equation
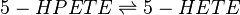
Rate equation
Enzyme Parameters
Kms
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 3.00E-01 | 
|
Human | Expression Vector: Bioimprited Enzyme - Selenosubtilisin
Enzyme: Glutathione Peroxidase pH: 7 Temperature: 37 |
[1] |
| 11.1 ± 2.90E-01 | 
|
Human | Expression Vector: E Coli
Enzyme: Wild Type Glutathione Peroxidase Enzyme (Se-hGSTZ1-1) pH: 7 Temperature: 37 |
[2] |
| 4.00E-03 | 
|
Rat | Expression Vector: Rat Liver Cells
Enzyme: Glutathione Peroxidase pH: 7.4 Temperature: 37 |
[3] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 3.19E-01 | 1.02E+03 | 2.54E+00 | 1.92E+00 |
Kmp
This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail here). As a result, no confidence interval factor or literature values were cited for this parameter.
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 3.15E-01 | 2.53E+00 | 1.92E+00 |
kcat
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 1170 ± 50 | 
|
Human | Expression Vector: Bioimprited Enzyme - Selenosubtilisin
Enzyme: Glutathione Peroxidase pH: 7 Temperature: 37 |
[1] |
| 24500 ± 150 | 
|
Human | Expression Vector: E Coli
Enzyme: Wild Type Glutathione Peroxidase Enzyme (Se-hGSTZ1-1) pH: 7 Temperature: 37 |
[2] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 2.44E+04 | 4.20E+00 | 1.11E+01 | 9.75E-01 |
Enzyme concentration
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 459 | 
|
Human | Expression Vector: Skin
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
[4] |
| 307 | 
|
Human | Expression Vector: Lung
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
[4] |
| 282 | 
|
Human | Expression Vector: Esophagus
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
[4] |
| 204 | 
|
Human | Expression Vector: Gut
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
[5] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 3.14E+02 | 1.38E+00 | 5.85E+00 | 3.09E-01 |
Keq
| Gibbs Free Energy Change | Units | Species | Notes | Reference |
|---|---|---|---|---|
| (-26.941177) | kcal/mol | Not stated | Estimated
Enzyme: PHGPx Substrate:a hydroperoxy-fatty-acyl-[lipid] Product: a hydroxy-fatty-acyl-[lipid] pH: 7.3 ionic strength: 0.25 |
[6] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 5.90E+19 | 1.00E+01 | 4.63E+01 | 8.90E-01 |
References
- ↑ 1.0 1.1 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein. Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.
- ↑ 2.0 2.1 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases. Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.
- ↑ Hiratsuka A. "Subunit Ya-specific glutathione peroxidase activity toward cholesterol 7-hydroperoxides of glutathione S-transferases in cytosols from rat liver and skin. J Biol Chem. 1997 Feb 21;272(8):4763-9.
- ↑ 4.0 4.1 4.2 M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471