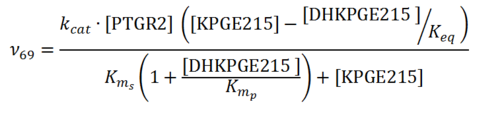Difference between revisions of "Transformation of 15-Keto-PGE2 to 13,14-Dihydro-15-Keto-PGE2"
(→Reaction) |
(→Parameters) |
||
| (23 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
[[Welcome to the In-Silico Model of Cutaneous Lipids Wiki | Return to overview]] | [[Welcome to the In-Silico Model of Cutaneous Lipids Wiki | Return to overview]] | ||
| − | + | The second step of the catabolic pathway of prostanoids is the reduction of the conjugated α, β-unsaturated double bond at C13, by 13, 15-ketoprostglandin reductase, also known as prostaglandin reductase. There are two isoforms of this protein, prostaglandin reductase 1 (PTGR-1) and prostaglandin reductase 2 (PTGR-2). Prostaglandin reductase 1 (PTGR-1) can accept a wide variety of prostaglandins as substrates. Prostaglandin reductase 2 (PTGR-2) has the highest affinity for 15-keto-PGE2, but also accepts a wide variety of prostaglandins as a substrate <ref>Wu, Yu-Hauh Ko, Tzu-Ping Guo, Rey-Ting Hu, Su-Ming Chuang, Lee-Ming Wang, Andrew H J., ''Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2'',Structure (2008), 16, 1714-1723.</ref>. | |
| − | |||
== Reaction == | == Reaction == | ||
| Line 8: | Line 7: | ||
==Chemical equation== | ==Chemical equation== | ||
| − | <center><math> 15-Keto-PGE2 \rightleftharpoons 13,14-Dihydro-15-Keto-PGE2</math></center> | + | <center><math> 15-Keto-PGE2 + NAD(P)+ \rightleftharpoons 13,14-Dihydro-15-Keto-PGE2 + NAD(P)H</math></center> |
== Rate equation == | == Rate equation == | ||
| + | [[File:R69.PNG|center|500px]] | ||
| + | |||
| + | == Parameters == | ||
| + | ===K<sub>ms</sub>=== | ||
| + | |||
| + | {|class="wikitable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 0.01121 ± 0.00014 | ||
| + | | mM | ||
| + | | Human | ||
| + | |Method: In vitro | ||
| + | Organism: Human | ||
| + | Expression vector: E.coli | ||
| + | Enzyme: PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 ◦C | ||
| + | Substrate: 15-Keto-PGE2 | ||
| + | |||
| + | "For determining the KM and Vmax values for | ||
| + | NADPH, 15-keto-PGE2 at a final concentration of 200 mM was used with different | ||
| + | concentrations of NADPH (0–60 mM)." | ||
| + | |512 | ||
| + | | <ref name="Wu2008"> [https://www.ncbi.nlm.nih.gov/pubmed/19000823 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.]</ref> | ||
| + | |- | ||
| + | |0.01587 ± 0.00171 | ||
| + | |mM | ||
| + | |Human | ||
| + | |Method: In vitro | ||
| + | Organism: Human | ||
| + | Expression vector: E.coli | ||
| + | Enzyme: e PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 ◦C | ||
| + | Substrate: NADPH | ||
| + | |||
| + | "For determining the KM and Vmax values for | ||
| + | NADPH, 15-keto-PGE2 at a final concentration of 200 mM was used with different | ||
| + | concentrations of NADPH (0–60 mM)." | ||
| + | |512 | ||
| + | | <ref name="Wu2008"> [https://www.ncbi.nlm.nih.gov/pubmed/19000823 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PTGR-2 Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.20E-02 || 4.92E+00 || -3.97E+00 || 6.71E-01 | ||
| + | |} | ||
| + | |||
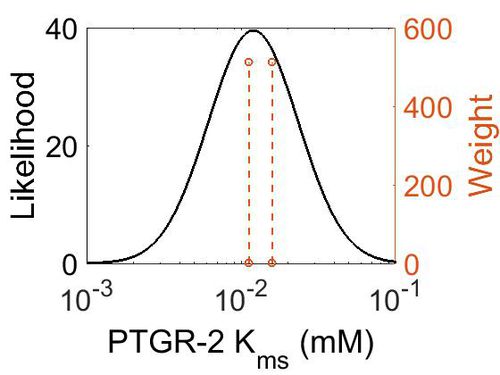
| + | [[Image:77.jpg|none|thumb|500px|The estimated probability distribution for PTGR-2 Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | ===K<sub>mp</sub>=== | ||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PTGR-2 Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.19E-02 || -3.97E+00 || 6.81E-01 | ||
| + | |- | ||
| + | | | ||
| + | |} | ||
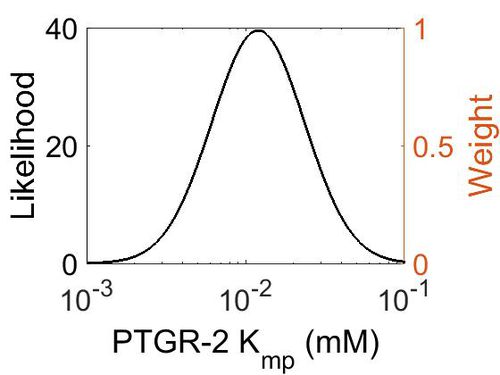
| + | [[Image:78.jpg|none|thumb|500px|The estimated probability distribution for PTGR-2 Kmp. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| − | == | + | ===k<sub>cat</sub>=== |
| + | Note: Turnover values from EC 1.3.1.48 - 13,14-dehydro-15-oxoprostaglandin 13-reductase | ||
| + | |||
| + | {|class="wikitable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 11.4 ± 0.9 | ||
| + | | min-1 | ||
| + | | Mouse | ||
| + | |Method: In vitro | ||
| + | Organism: Mouse | ||
| + | Expression vector: | ||
| + | Enzyme: 13,14-dehydro-15-oxoprostaglandin 13-reductase | ||
| + | pH: 7.4 | ||
| + | Temperature: 37 ◦C | ||
| + | Substrate: 15-Ketoprostaglandin E2 | ||
| + | |384 | ||
| + | |<ref name="Wu2008"> [https://www.ncbi.nlm.nih.gov/pubmed/19000823 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PTGR-2 kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.14E+01 || 9.86E+00 || 3.22E+00 || 8.87E-01 | ||
| + | |} | ||
| + | |||
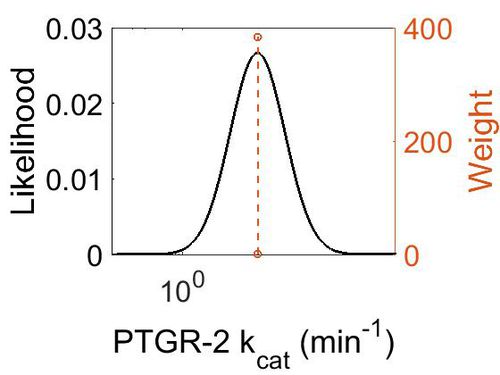
| + | [[Image:79.jpg|none|thumb|500px|The estimated probability distribution for PTGR-2 kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | ===Enzyme concentration === | ||
| + | |||
| + | To convert the enzyme concentration from ppm to mM, the following [[Common equations#Enzyme concentration (mM)|equation]] was used. | ||
| + | |||
| + | {|class="wikitable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 162 | ||
| + | | ppm | ||
| + | | Human | ||
| + | |Expression Vector: Skin | ||
| + | Enzyme: PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |2048 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the | ||
| + | human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |80.9 | ||
| + | |ppm | ||
| + | |Human | ||
| + | |Expression Vector: Skin | ||
| + | Enzyme: PTGR2 | ||
| + | pH: Unknown | ||
| + | Temperature: Unknown | ||
| + | |2048 | ||
| + | |Unknown | ||
| + | |- | ||
| + | |74.1 | ||
| + | |ppm | ||
| + | |Human | ||
| + | |Expression Vector: Oral Cavity | ||
| + | Enzyme: PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the | ||
| + | human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PTGR-2 concentration distribution | ||
| + | ! Mode (ppm) !! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 8.65E+01 || 4.79E-04|| 1.45E+00 || 4.58E+00 || 3.47E-01 | ||
| + | |} | ||
| + | |||
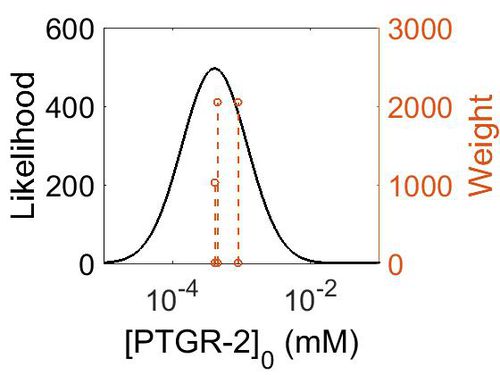
| + | [[Image:168.jpg|none|thumb|500px|The estimated probability distribution for PTGR-2 concentration. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | ===K<sub>eq</sub>=== | ||
| + | {|class="wikitable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | ! Gibbs Free energy | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 3.6006165 | ||
| + | | kcal/mol | ||
| + | |Unspecified | ||
| + | |Calculations with a Gaussian98 suite of programs | ||
| + | Enzyme: COX (Unspecific) | ||
| + | Substrate: Arachidonate | ||
| + | Temperature: 298.15 K | ||
| + | Pressure: 1 bar | ||
| + | |64 | ||
| + | |<ref name="Silva2003”>[http://link.springer.com/article/10.1007/s00214-003-0476-9 P. Silva, "A theoretical study of radical-only and combined radical/carbocationic mechanisms of arachidonic acid cyclooxygenation by prostaglandin H synthase" Theor Chem Acc (2003) 110: 345]</ref> | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PTGR-2 Keq distribution | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 2.28E-03 || 1.00E+01 || -5.29E+00 || 8.91E-01 | ||
| + | |} | ||
| + | |||
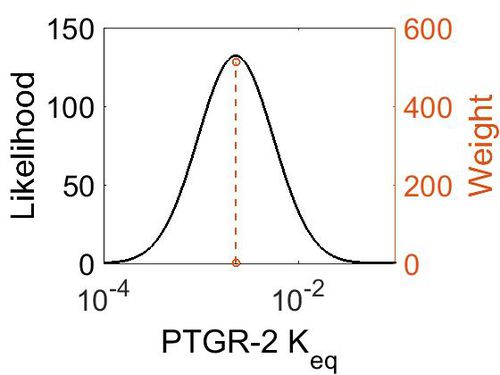
| + | [[Image:80.jpg|none|thumb|500px|The estimated probability distribution for PTGR-2 Keq. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | ===Misch=== | ||
{|class="wikitable" | {|class="wikitable" | ||
| Line 31: | Line 223: | ||
| <ref name="Bothwell1982”>[http://jpet.aspetjournals.org/content/220/2/229.long W. Bothwell, "A radioimmunoassay for the unstable pulmonary metabolites of prostaglandin E1 and E2: an indirect index of their in vivo disposition and pharmacokinetics" Journal of Pharmacology and Experimental Therapeutics February 1982, 220 (2) 229-235]</ref> | | <ref name="Bothwell1982”>[http://jpet.aspetjournals.org/content/220/2/229.long W. Bothwell, "A radioimmunoassay for the unstable pulmonary metabolites of prostaglandin E1 and E2: an indirect index of their in vivo disposition and pharmacokinetics" Journal of Pharmacology and Experimental Therapeutics February 1982, 220 (2) 229-235]</ref> | ||
|} | |} | ||
| + | |||
| + | {|class="wikitable" | ||
| + | |+ style="text-align: left;" | Vmax | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 159.23 ± 0.71 | ||
| + | | nmol min-1 mg-1 | ||
| + | | Human | ||
| + | |Method: In vitro | ||
| + | Organism: Human | ||
| + | Expression vector: E.coli | ||
| + | Enzyme: PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 ◦C | ||
| + | Substrate: 15-Keto-PGE2 | ||
| + | | <ref name="Wu2008"> [https://www.ncbi.nlm.nih.gov/pubmed/19000823 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.]</ref> | ||
| + | |- | ||
| + | | 66.73 ± 1.36 | ||
| + | | nmol min-1 mg-1 | ||
| + | | Human | ||
| + | |Method: In vitro | ||
| + | Organism: Human | ||
| + | Expression vector: E.coli | ||
| + | Enzyme: PTGR2 | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 ◦C | ||
| + | Substrate: NADPH | ||
| + | | <ref name="Wu2008"> [https://www.ncbi.nlm.nih.gov/pubmed/19000823 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | == References == | ||
| + | <references/> | ||
Latest revision as of 11:09, 2 November 2019
The second step of the catabolic pathway of prostanoids is the reduction of the conjugated α, β-unsaturated double bond at C13, by 13, 15-ketoprostglandin reductase, also known as prostaglandin reductase. There are two isoforms of this protein, prostaglandin reductase 1 (PTGR-1) and prostaglandin reductase 2 (PTGR-2). Prostaglandin reductase 1 (PTGR-1) can accept a wide variety of prostaglandins as substrates. Prostaglandin reductase 2 (PTGR-2) has the highest affinity for 15-keto-PGE2, but also accepts a wide variety of prostaglandins as a substrate [1].
Contents
Reaction
Chemical equation
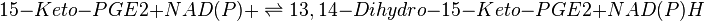
Rate equation
Parameters
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 0.01121 ± 0.00014 | mM | Human | Method: In vitro
Organism: Human Expression vector: E.coli Enzyme: PTGR2 pH: 7.5 Temperature: 37 ◦C Substrate: 15-Keto-PGE2 "For determining the KM and Vmax values for NADPH, 15-keto-PGE2 at a final concentration of 200 mM was used with different concentrations of NADPH (0–60 mM)." |
512 | [2] |
| 0.01587 ± 0.00171 | mM | Human | Method: In vitro
Organism: Human Expression vector: E.coli Enzyme: e PTGR2 pH: 7.5 Temperature: 37 ◦C Substrate: NADPH "For determining the KM and Vmax values for NADPH, 15-keto-PGE2 at a final concentration of 200 mM was used with different concentrations of NADPH (0–60 mM)." |
512 | [2] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.20E-02 | 4.92E+00 | -3.97E+00 | 6.71E-01 |
Kmp
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 1.19E-02 | -3.97E+00 | 6.81E-01 |
kcat
Note: Turnover values from EC 1.3.1.48 - 13,14-dehydro-15-oxoprostaglandin 13-reductase
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 11.4 ± 0.9 | min-1 | Mouse | Method: In vitro
Organism: Mouse Expression vector: Enzyme: 13,14-dehydro-15-oxoprostaglandin 13-reductase pH: 7.4 Temperature: 37 ◦C Substrate: 15-Ketoprostaglandin E2 |
384 | [2] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.14E+01 | 9.86E+00 | 3.22E+00 | 8.87E-01 |
Enzyme concentration
To convert the enzyme concentration from ppm to mM, the following equation was used.
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 162 | ppm | Human | Expression Vector: Skin
Enzyme: PTGR2 pH: 7.5 Temperature: 37 °C |
2048 | [3] |
| 80.9 | ppm | Human | Expression Vector: Skin
Enzyme: PTGR2 pH: Unknown Temperature: Unknown |
2048 | Unknown |
| 74.1 | ppm | Human | Expression Vector: Oral Cavity
Enzyme: PTGR2 pH: 7.5 Temperature: 37 °C |
1024 | [3] |
| Mode (ppm) | Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|---|
| 8.65E+01 | 4.79E-04 | 1.45E+00 | 4.58E+00 | 3.47E-01 |
Keq
| Gibbs Free energy | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 3.6006165 | kcal/mol | Unspecified | Calculations with a Gaussian98 suite of programs
Enzyme: COX (Unspecific) Substrate: Arachidonate Temperature: 298.15 K Pressure: 1 bar |
64 | [4] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 2.28E-03 | 1.00E+01 | -5.29E+00 | 8.91E-01 |
Misch
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 9.9 +/-0.2 | minutes | Dog | In vivo
Temperature:37 pH:7 |
[5] |
| Value | Units | Species | Notes | Reference |
|---|---|---|---|---|
| 159.23 ± 0.71 | nmol min-1 mg-1 | Human | Method: In vitro
Organism: Human Expression vector: E.coli Enzyme: PTGR2 pH: 7.5 Temperature: 37 ◦C Substrate: 15-Keto-PGE2 |
[2] |
| 66.73 ± 1.36 | nmol min-1 mg-1 | Human | Method: In vitro
Organism: Human Expression vector: E.coli Enzyme: PTGR2 pH: 7.5 Temperature: 37 ◦C Substrate: NADPH |
[2] |
References
- ↑ Wu, Yu-Hauh Ko, Tzu-Ping Guo, Rey-Ting Hu, Su-Ming Chuang, Lee-Ming Wang, Andrew H J., Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2,Structure (2008), 16, 1714-1723.
- ↑ 2.0 2.1 2.2 2.3 2.4 "Structural basis for catalytic and inhibitory mechanisms of human prostaglandin reductase PTGR2", Structure. 2008 Nov 12;16(11):1714-23. doi: 10.1016/j.str.2008.09.007.
- ↑ 3.0 3.1 [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587]
- ↑ P. Silva, "A theoretical study of radical-only and combined radical/carbocationic mechanisms of arachidonic acid cyclooxygenation by prostaglandin H synthase" Theor Chem Acc (2003) 110: 345
- ↑ W. Bothwell, "A radioimmunoassay for the unstable pulmonary metabolites of prostaglandin E1 and E2: an indirect index of their in vivo disposition and pharmacokinetics" Journal of Pharmacology and Experimental Therapeutics February 1982, 220 (2) 229-235