Difference between revisions of "Dynamics of bacterial colony growth"
| Line 1: | Line 1: | ||
For our simulations we assume that cells are maintained in the exponential phase with doubling time <math>\tau</math>. In our virtual colony, each individual cell grows exponentially in time until its division into two daughter cells, as per | For our simulations we assume that cells are maintained in the exponential phase with doubling time <math>\tau</math>. In our virtual colony, each individual cell grows exponentially in time until its division into two daughter cells, as per | ||
| − | <center><math>V_{c,i}(t)=V_{0}\cdot 2^{t/\tau_{i}}</math>,</center> | + | <center><math>V_{c,i}(t)=V_{0}\cdot 2^{t/ \tau_{i}}</math>,</center> |
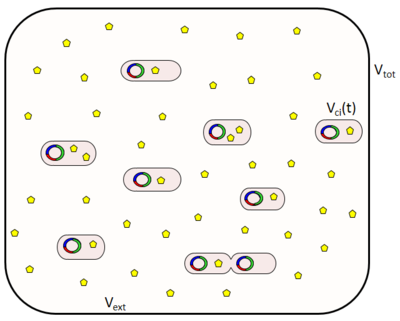
where <math>V_{0}</math> is the volume of a cell at the beginning of the cell cycle (same for all cells), <math>\tau_{i}</math> is the duration of the cell cycle of cell ''i'', and ''t'' is the time to the precedent division event. When <math>t = \tau_{i}</math> the cell i has doubled its volume and a new division takes place. At this timepoint the internal clocks and volumes of the daughter cells are reset to zero and <math>V_{0}</math> respectively. Moreover, when a cell divides, proteins, mRNAs and signalling molecules are binomially distributed <ref name="Rosenfeld2005"> [http://www.sciencemag.org/content/307/5717/1962.full.pdf Rosenfeld N, Young JW, Alon U, Swain PS, Elowitz MB. ''Gene regulation at the single-cell level.'' Sci (New York, N.Y.) 2005, 307(5717):1962–5.]</ref> between daughter cells and one copy of the DNA is given to each cell. The regulatory complexes bound to the DNA are detached prior to the distribution between daughter cells. The total volume of the culture <math>V_{tot}</math> is constant and is composed by the volume of the total cells and the volume of the medium, as per the equation <math>V_{tot} = V_{ext} + \displaystyle\sum_{j=1}^{N}V_{c,j}(t)</math>. | where <math>V_{0}</math> is the volume of a cell at the beginning of the cell cycle (same for all cells), <math>\tau_{i}</math> is the duration of the cell cycle of cell ''i'', and ''t'' is the time to the precedent division event. When <math>t = \tau_{i}</math> the cell i has doubled its volume and a new division takes place. At this timepoint the internal clocks and volumes of the daughter cells are reset to zero and <math>V_{0}</math> respectively. Moreover, when a cell divides, proteins, mRNAs and signalling molecules are binomially distributed <ref name="Rosenfeld2005"> [http://www.sciencemag.org/content/307/5717/1962.full.pdf Rosenfeld N, Young JW, Alon U, Swain PS, Elowitz MB. ''Gene regulation at the single-cell level.'' Sci (New York, N.Y.) 2005, 307(5717):1962–5.]</ref> between daughter cells and one copy of the DNA is given to each cell. The regulatory complexes bound to the DNA are detached prior to the distribution between daughter cells. The total volume of the culture <math>V_{tot}</math> is constant and is composed by the volume of the total cells and the volume of the medium, as per the equation <math>V_{tot} = V_{ext} + \displaystyle\sum_{j=1}^{N}V_{c,j}(t)</math>. | ||
| Line 11: | Line 11: | ||
where <math>\tau</math> and <math>\tilde{\tau}</math> correspond to the deterministic and stochastic parameters of the cell cycle duration respectively, and <math>\lambda \in [0, 1] </math> is a parameter that weights their relative importance.The stochastic component accounts for the period of time between events described by a Poissonian process according to an exponential distribution, | where <math>\tau</math> and <math>\tilde{\tau}</math> correspond to the deterministic and stochastic parameters of the cell cycle duration respectively, and <math>\lambda \in [0, 1] </math> is a parameter that weights their relative importance.The stochastic component accounts for the period of time between events described by a Poissonian process according to an exponential distribution, | ||
<center><math>\phi(\tilde{\tau})=\frac{e^{-\frac{\tilde{\tau}}{\tau}}}{\tau}</math>.</center> | <center><math>\phi(\tilde{\tau})=\frac{e^{-\frac{\tilde{\tau}}{\tau}}}{\tau}</math>.</center> | ||
| − | In this way, variability is enabled from cell to cell with regards to the duration of the cell cycle, yet a minimum cell cycle duration <math>\lambda\cdot \tau</math> is set. Therefore, the average duration and standard deviation of the cell cycle are <math>\tau</math> and <math>(1 | + | In this way, variability is enabled from cell to cell with regards to the duration of the cell cycle, yet a minimum cell cycle duration <math>\lambda\cdot \tau</math> is set. Therefore, the average duration and standard deviation of the cell cycle are <math>\tau</math> and <math>(1 - \lambda) \cdot \tau</math> respectively. <ref name="Weber2013"> [http://www.biomedcentral.com/content/pdf/1752-0509-7-6.pdf M. Weber and J. Buceta. ''Dynamics of the quorum sensing switch: Stochastic and non-stationary effects.'' BMC Systems Biology 2013,(7):6]</ref> |
The values of the cell growth parameters that are used in the model are summarized in the following table: | The values of the cell growth parameters that are used in the model are summarized in the following table: | ||
| Line 23: | Line 23: | ||
|<math>\tau</math> | |<math>\tau</math> | ||
|Cell cycle duration (doubling time) | |Cell cycle duration (doubling time) | ||
| − | | <math> 64. | + | | <math> 64.8-297.06 min</math> |
| The doubling time values are the ones reported in the section above. | | The doubling time values are the ones reported in the section above. | ||
|- | |- | ||
Latest revision as of 14:46, 29 January 2019
For our simulations we assume that cells are maintained in the exponential phase with doubling time 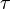 . In our virtual colony, each individual cell grows exponentially in time until its division into two daughter cells, as per
. In our virtual colony, each individual cell grows exponentially in time until its division into two daughter cells, as per
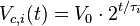 ,
,where 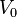 is the volume of a cell at the beginning of the cell cycle (same for all cells),
is the volume of a cell at the beginning of the cell cycle (same for all cells), 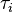 is the duration of the cell cycle of cell i, and t is the time to the precedent division event. When
is the duration of the cell cycle of cell i, and t is the time to the precedent division event. When 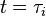 the cell i has doubled its volume and a new division takes place. At this timepoint the internal clocks and volumes of the daughter cells are reset to zero and
the cell i has doubled its volume and a new division takes place. At this timepoint the internal clocks and volumes of the daughter cells are reset to zero and 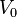 respectively. Moreover, when a cell divides, proteins, mRNAs and signalling molecules are binomially distributed [1] between daughter cells and one copy of the DNA is given to each cell. The regulatory complexes bound to the DNA are detached prior to the distribution between daughter cells. The total volume of the culture
respectively. Moreover, when a cell divides, proteins, mRNAs and signalling molecules are binomially distributed [1] between daughter cells and one copy of the DNA is given to each cell. The regulatory complexes bound to the DNA are detached prior to the distribution between daughter cells. The total volume of the culture 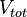 is constant and is composed by the volume of the total cells and the volume of the medium, as per the equation
is constant and is composed by the volume of the total cells and the volume of the medium, as per the equation 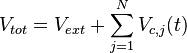 .
.
With regards to the effect of the cell volume of individual cells on the diffusion rate of the autoinducer, the coefficient  which contributes to the diffusion of the external autoinducer into the cells, is described by the equation
which contributes to the diffusion of the external autoinducer into the cells, is described by the equation
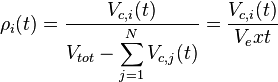 .
.The duration of the cell cycle, 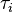 , is different for each cell and is set independently after a division according to the stochastic rule [2]
, is different for each cell and is set independently after a division according to the stochastic rule [2]
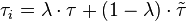 ,
,where 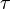 and
and 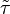 correspond to the deterministic and stochastic parameters of the cell cycle duration respectively, and
correspond to the deterministic and stochastic parameters of the cell cycle duration respectively, and ![\lambda \in [0, 1]](/wiki/images/math/a/d/a/ada29722c713d4d8a571d9a4ce353ae6.png) is a parameter that weights their relative importance.The stochastic component accounts for the period of time between events described by a Poissonian process according to an exponential distribution,
is a parameter that weights their relative importance.The stochastic component accounts for the period of time between events described by a Poissonian process according to an exponential distribution,
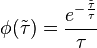 .
.In this way, variability is enabled from cell to cell with regards to the duration of the cell cycle, yet a minimum cell cycle duration 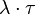 is set. Therefore, the average duration and standard deviation of the cell cycle are
is set. Therefore, the average duration and standard deviation of the cell cycle are 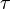 and
and 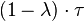 respectively. [3]
respectively. [3]
The values of the cell growth parameters that are used in the model are summarized in the following table:
| Name | Description | Value | Remarks-Reference |
|---|---|---|---|
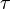
|
Cell cycle duration (doubling time) | 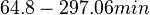
|
The doubling time values are the ones reported in the section above. |
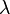
|
relative weight between the det.-stoch.
parameters of the cell cycle |
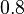 [1][4] [1][4]
|
Single cell level gene regulation studies in E. coli showed a difference of ~0.8 between single cell cycle duration and the mean cell cycle duration, due to periodic oscillations. |
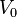
|
cell volume at the beginning of cell cycle | 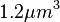 [5] [5]
|
The cell volume is derived by the publication of R.A. Cox (ref. 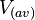 in the figure in Parameters section) where he reported the average cell volume for S. coelicolor A3(2) grown at 30 oC under different growth rates, between in the figure in Parameters section) where he reported the average cell volume for S. coelicolor A3(2) grown at 30 oC under different growth rates, between 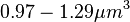
|
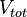
|
total cell culture volume | 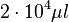
|
References
- ↑ 1.0 1.1 Rosenfeld N, Young JW, Alon U, Swain PS, Elowitz MB. Gene regulation at the single-cell level. Sci (New York, N.Y.) 2005, 307(5717):1962–5.
- ↑ Canela-Xandri O, Sagues F, Buceta J. Interplay between intrinsic noise and the stochasticity of the cell cycle in bacterial colonies. Biophys J 2010, 98(11):2459–68.
- ↑ M. Weber and J. Buceta. Dynamics of the quorum sensing switch: Stochastic and non-stationary effects. BMC Systems Biology 2013,(7):6
- ↑ Reshes G, Vanounou S, Fishov I, Feingold M: Timing the start of division in E. coli: a single-cell study. Phys Biol 2008, 5(4):046001.
- ↑ Cox RA. Quantitative relationships for specific growth rates and macromolecular compositions of Mycobacterium tuberculosis, Streptomyces coelicolor A3(2) and Escherichia coli B/r: an integrative theoretical approach. Microbiology. 2004 May;150(Pt 5):1413-26.