Difference between revisions of "Degradation of A-R2"
(→Parameters with uncertainty) |
(→Parameters with uncertainty) |
||
| Line 38: | Line 38: | ||
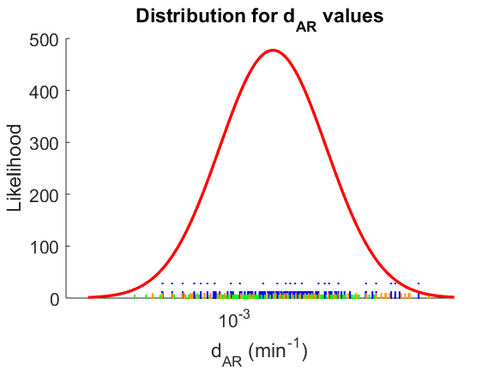
The probability distribution for the parameter, adjusted accordingly in order to reflect the above values, is the following: | The probability distribution for the parameter, adjusted accordingly in order to reflect the above values, is the following: | ||
| − | [[Image: | + | [[Image:DAR.png|500px]] |
The location and scale parameters of the distribution are: | The location and scale parameters of the distribution are: | ||
Revision as of 21:45, 28 September 2015
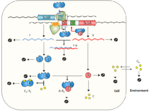
The ScbR-ScbA protein complex (AR2) degrades.
Contents
Chemical equation
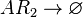
Rate equation
![r= d_{AR}\cdot[AR_{2}]](/wiki/images/math/a/1/8/a182be77e40bd08d4d2dbcd1f3bc70f6.png)
Parameters
The parameter of this reaction is the degradation rate of AR2 (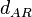 ).
).
| Name | Value | Units | Origin | Remarks |
|---|---|---|---|---|
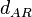
|
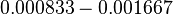 [1] [1]
|

|
Degradation rates of proteins in S. coelicolor
based on radioactive decay |
Parameters with uncertainty
The most plausible parameter value for the 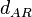 is decided to be
is decided to be 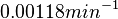 and the confidence interval
and the confidence interval 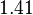 . This means that the mode of the PDF is 0.00118 and the range where 95% of the values are found is between
. This means that the mode of the PDF is 0.00118 and the range where 95% of the values are found is between 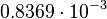 and
and 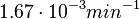 .
.
The probability distribution for the parameter, adjusted accordingly in order to reflect the above values, is the following:
The location and scale parameters of the distribution are:
| Parameter | μ | σ |
|---|---|---|
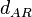
|
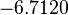
|
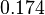
|
References
- ↑ Trötschel C, Albaum SP, Poetsch A. Proteome turnover in bacteria: current status for Corynebacterium glutamicum and related bacteria. Microbial biotechnology. 2013;6(6):708-719.