Difference between revisions of "DXS"
Aliah.hawari (talk | contribs) (→DXS parameters) |
Aliah.hawari (talk | contribs) (→DXS parameters) |
||
| Line 20: | Line 20: | ||
== DXS parameters == | == DXS parameters == | ||
| + | Parameterisation for DXS is done using our PROMENADE pipeline where values measured in '' E. coli '' and close to ''in vivo'' conditions are weighted higher. More information on how I parameterize my model using the pipeline can be found here. | ||
| + | |||
[[File:DXS_Kcat.png | left |600px ]] | [[File:DXS_Kcat.png | left |600px ]] | ||
Latest revision as of 15:22, 23 March 2017
You can go back to main page of the kinetic model here.
The DXS reaction (EC 2.2.1.7)
Deoxyxylulose-5-phosphate synthase (DXS) catalyses the production of 1-deoxy-D-xylulose 5-phosphate (DXP) from pyruvate and glyceraldehyde 3-phosphate (G3P). This reaction is the first step in the MEP pathway.
Modelling DXS
In the kinetic model, the DXS reaction is modelled with reversible Michaelis-Menten using the Hanekom [1] bi-bi random order generic equation. In total, this reaction requires five kinetic parameters (Kms for all substrates and products, and a forward Kcat) and one thermodynamic parameter (Equilibrium constant, Keq).
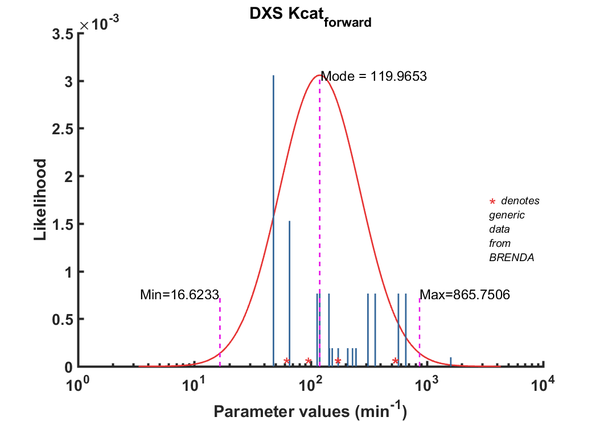
DXS parameters
Parameterisation for DXS is done using our PROMENADE pipeline where values measured in E. coli and close to in vivo conditions are weighted higher. More information on how I parameterize my model using the pipeline can be found here.
| Parameter | Direction | Substrate | Value | Unit | Weight | Description | Reference |
|---|---|---|---|---|---|---|---|
| Kcat | Forward | DXS | 229.7 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [2] |
| Kcat | Forward | DXS | 153.6 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [2] |
| Kcat | Forward | DXS | 145.5 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [2] |
| Kcat | Forward | DXS | 209 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [2] |
| Kcat | Forward | DXS | 173 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [2] |
| Kcat | Forward | DXS | 246 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [2] |
| Kcat | Forward | GAP | 48 | 1/min | 256 | Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. | [3] |
| Kcat | Forward | GAP | 66 | 1/min | 128 | DXPS2; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | [4] |
| Kcat | Forward | GAP | 114 | 1/min | 8 | from R. capsulatus, pH 7.4, 37C | [5] |
| Kcat | Forward | GAP | 660 | 1/min | 64 | from Plasmodium, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | [6] |
| Kcat | Forward | GAP | 1608 | 1/min | 8 | dxs11 from Agrobacterium tumefaciens, pH8.0, 37¡C, expressed in E. coli | [7] |
| Kcat | Forward | GAP | 120 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Kcat | Forward | GAP | 120 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Kcat | Forward | GAP | 360 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Kcat | Forward | Pyruvate | 570 | 1/min | 64 | from Plasmodium, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | [6] |
| Kcat | Forward | Pyruvate | 144 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Kcat | Forward | Pyruvate | 114 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Kcat | Forward | Pyruvate | 312 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Parameter | Direction | Substrate | Value | Unit | Weight | Description | Reference |
|---|---|---|---|---|---|---|---|
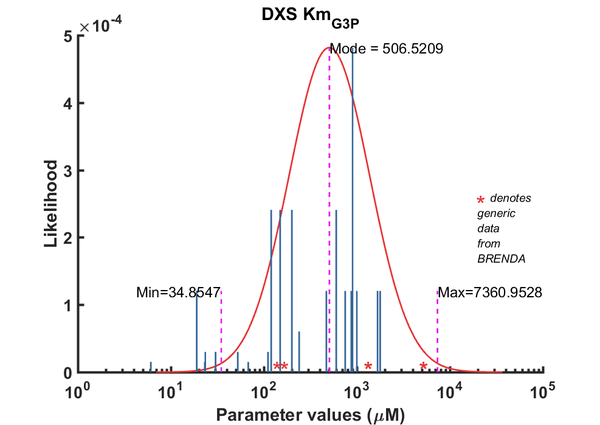
| Km | Forward | G3P | 6.1 | µM | 8 | pH 7.9, 37C (optimal), Mycobacterium tuberculosis, Km_G3P 6.1 microM, Km_Pyr 40 microM | Bailey2002, Br658834 |
| Km | Forward | G3P | 1670 | µM | 64 | from Vitis vinifera L. ; expressed in E. coli DH5a, pH 8.0; 37C; VvDXS N284 | Battilana2011, Br720131 |
| Km | Forward | G3P | 1780 | µM | 64 | from Vitis vinifera L. ; expressed in E. coli DH5a, pH 8.0; 37C; VvDXS N284 | Battilana2011, Br720131 |
| Km | Forward | G3P | 900 | µM | 256 | Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. | Boronat1999 |
| Km | Forward | G3P | 750 | µM | 64 | from Capsicum anuum; Km_pyruvate: 500 microM; Km_GAP: 750 microM | Bouvier1998 |
| Km | Forward | G3P | 30.3 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | [2] |
| Km | Forward | G3P | 23.5 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min. Kinetics were measured in different buffers (buffer optimization study). All buffers ph*, 37C | [2] |
| Km | Forward | G3P | 52.5 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | [2] |
| Km | Forward | G3P | 600 | µM | 128 | DXPS1; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | [4] |
| Km | Forward | G3P | 200 | µM | 128 | DXPS2; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | [4] |
| Km | Forward | G3P | 68 | µM | 8 | from Rhodobacter capsulatus, pH7.4, 37C, Km_GAP: 0.068 +/- 0.001 mM, Km_pyr: 0.44 +/- 0.05 mM. | [5] |
| Km | Forward | G3P | 111 | µM | 16 | from Grey poplar leaves, pH 8.6; 45 C; | Ghirardo2010 |
| Km | Forward | G3P | 150 | µM | 128 | from Rhodobacter capsulatus, pH7.4, 37¡C; expressed in E. coli; recombinant isoenzyme-A; Km_pyruvate: 610 +/- 50 microM; Km_GAP: 150 +/- 10 microM; Vmax = 1.9 +/- 0.1 micrommol/min/mg; | Hahn2001, Br395816 |
| Km | Forward | G3P | 120 | µM | 128 | from Rhodobacter capsulatus, pH7.4, 37¡C; expressed in E. coli; recombinant isoenzyme-A; Km_pyruvate: 610 +/- 50 microM; Km_GAP: 150 +/- 10 microM; Vmax = 1.9 +/- 0.1 micrommol/min/mg; | Hahn2001, Br395816 |
| Km | Forward | G3P | 19 | µM | 64 | from Plasmodium, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | [6] |
| Km | Forward | G3P | 120 | µM | 8 | from Streptomyces sp. strain CL190, expressed in E. coli. pH 9.0, 42-44C. Table 1. Km_pyruvate: 65microM, Km_GAP: 120 microM | Kuzuyama2000, Br395812 |
| Km | Forward | G3P | 240 | µM | 32 | from E. coli. pH 7.5-8.0.0, 42-44C. Km_pyruvate: 65microM, Km_GAP: 120 microM. See Table 1. Measured to compare with DXS from streptomyces. Purified in Kuzuyama1998 study. | Kuzuyama2000, Br395812 |
| Km | Forward | G3P | 23.2 | µM | 8 | dxs11 from Agrobacterium tumefaciens, pH8.0, 37¡C, expressed in E. coli | Lee2007, Br674982 |
| Km | Forward | G3P | 470 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Km | Forward | G3P | 870 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Km | Forward | G3P | 1000 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | [8] |
| Parameter | Direction | Substrate | Value | Unit | Weight | Description | Reference |
|---|---|---|---|---|---|---|---|
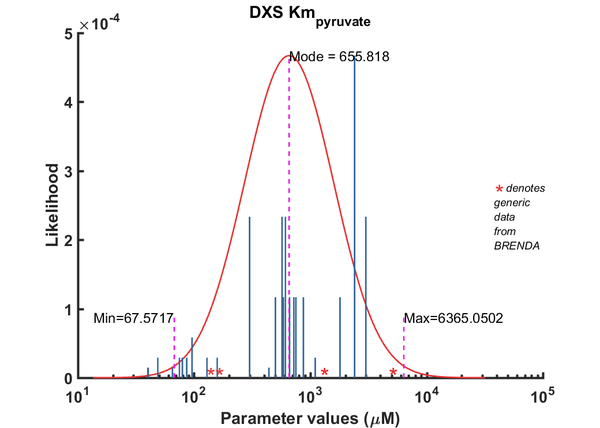
| Km | Forward | Pyruvate | 40 | µM | 8 | pH 7.9, 37C (optimal), Mycobacterium tuberculosis, Km_G3P 6.1 microM, Km_Pyr 40 microM | Bailey2002, Br658834 |
| Km | Forward | Pyruvate | 585.2 | µM | 64 | from Vitis vinifera L. ; expressed in E. coli DH5a, pH 8.0; 37C; VvDXS N284 & K284 (allelic variants) | Battilana2011, Br720131 |
| Km | Forward | Pyruvate | 661.2 | µM | 64 | from Vitis vinifera L. ; expressed in E. coli DH5a, pH 8.0; 37C; VvDXS N284 | Battilana2011, Br720131 |
| Km | Forward | Pyruvate | 2400 | µM | 256 | Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. | Boronat1999 |
| Km | Forward | Pyruvate | 500 | µM | 64 | from Capsicum anuum; pH 7.6, 30¡C. Km_pyruvate: 500 microM; Km_GAP: 750 microM | Bouvier1998 |
| Km | Forward | Pyruvate | 79.3 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 48.7 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 86.3 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 74.7 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 129 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 1100 | µM | 16 | from E. coli wild type DXS, Optimal buffer: 100 mM HEPES, pH 8, 37C ; Km_GAP:23.5 +/- 1.7 microM; Km_pyruvate: 48.7 +/- 8.0 microM, kcat: 153.6 +/- 6.8 1/min | Brammer2011, Br719975 |
| Km | Forward | Pyruvate | 300 | µM | 128 | DXPS1; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | Cane2001 |
| Km | Forward | Pyruvate | 570 | µM | 128 | DXPS2; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | Cane2001 |
| Km | Forward | Pyruvate | 440 | µM | 8 | from Rhodobacter capsulatus, pH7.4, 37C, Km_GAP: 0.068 +/- 0.001 mM, Km_pyr: 0.44 +/- 0.05 mM. | Eubanks2003, Br657953 |
| Km | Forward | Pyruvate | 158 | µM | 16 | from Grey poplar leaves, pH 8.6; 45 C; | Ghirardo2010 |
| Km | Forward | Pyruvate | 610 | µM | 128 | from Rhodobacter capsulatus, pH7.4, 37¡C; expressed in E. coli; recombinant isoenzyme-A; Km_pyruvate: 610 +/- 50 microM; Km_GAP: 150 +/- 10 microM; Vmax = 1.9 +/- 0.1 micrommol/min/mg; | Hahn2001, Br395816 |
| Km | Forward | Pyruvate | 3000 | µM | 128 | from Rhodobacter capsulatus, pH7.4, 37¡C; expressed in E. coli; recombinant isoenzyme-A; Km_pyruvate: 610 +/- 50 microM; Km_GAP: 150 +/- 10 microM; Vmax = 1.9 +/- 0.1 micrommol/min/mg; | Hahn2001, Br395816 |
| Km | Forward | Pyruvate | 870 | µM | 64 | from Plasmodium vivax, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | Handa2013, Br719510 |
| Km | Forward | Pyruvate | 65 | µM | 8 | from Streptomyces sp. strain CL190, expressed in E. coli. pH 9.0, 42-44C. Km_pyruvate: 65microM, Km_GAP: 120 microM | Kuzuyama2000, Br395812 |
| Km | Forward | Pyruvate | 96 | µM | 32 | from E. coli. pH 7.5-8.0., 42-44C. Km_pyruvate: 65microM, Km_GAP: 120 microM. See Table 1. Measured to compare with DXS from streptomyces. Purified in Kuzuyama1998 study. | Kuzuyama2000, Br395812 |
| Km | Forward | Pyruvate | 40.3 | µM | 8 | dxs11 from Agrobacterium tumefaciens, pH8.0, 37¡C, expressed in E. coli | Lee2007, Br674982 |
| Km | Forward | Pyruvate | 1800 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Km | Forward | Pyruvate | 720 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Km | Forward | Pyruvate | 750 | µM | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
References
- ↑ Hanekom, A. J. 2006. "Generic kinetic equations for modelling multisubstrate reactions in computational systems biology", MSc Thesis submitted at the University of Stellenbosch
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 2.7 2.8 Brammer, L.A. 2011 "1-FDeoxy-D-xylulose 5-phosphate synthase catalyzes a novel random sequential mechanism", JBioChem, 283(42):36522-36531.
- ↑ Brammer, L.A. 2011 "1-Deoxy-D-xylulose 5-phosphate synthase catalyzes a novel random sequential mechanism", JBioChem, 283(42):36522-36531.
- ↑ 4.0 4.1 4.2 Cane, D.E., et. al.. 2001 "Molecular cloning, expression and characterization of the first three genes in the mevalonate-independent isoprenoid pathway in Streptomyces coelicolor ", Bioorganic & Medicinal Chemistry, 9:1467-1477.
- ↑ 5.0 5.1 Eubanks, L.M. & Poulter, C.D. 2003. "Rhodobacter capsulatus 1-Deoxy-D-xylulose 5-Phosphate Synthase: Steady-State Kinetics and Substrate Binding† ", Biochemistry, 42:1140-1149.
- ↑ 6.0 6.1 6.2 Handa, S. et. al. 2013. "Production of recombinant 1-deoxy-d-xylulose-5-phosphate synthase from Plasmodium vivax in Escherichia coli", Biochemistry, 3:124-129. Cite error: Invalid
<ref>tag; name "Handa2013" defined multiple times with different content - ↑ Lee, J. et. al. 2007. "Cloning and characterization of the dxs gene, encoding 1-deoxy-d-xylulose 5-phosphate synthase from Agrobacterium tumefaciens, and its overexpression in Agrobacterium tumefaciens", JBiotech, 128:555-566.
- ↑ 8.0 8.1 8.2 8.3 8.4 8.5 8.6 8.7 8.8 Matsushima, D. et. al2012. "The single cellular green microalga Botryococcus braunii, race B possesses three distinct 1-deoxy-d-xylulose 5-phosphate synthases", PlantSci, 185-186:309-320.
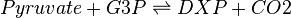
![V_\mathrm{DXS}= \cfrac{Kcat_\mathrm{forward} \bullet [DXS] \bullet \left( \cfrac{[Pyr]}{Km_\mathrm{DXS}} \right) \bullet \left( \cfrac{[G3P]}{Km_\mathrm{g3p}} \right) \bullet \left( 1 - \cfrac{\left( \cfrac{[DXP]\bullet[CO2]}{[Pyr]\bullet[G3P]} \right)}{K_\mathrm{eq}} \right)} {\left( 1 + \cfrac {[Pyr]}{Km_\mathrm{pyr}} + \cfrac{[CO2]}{Km_\mathrm{co2}}\right) \bullet \left( 1 + \cfrac{[G3P]}{Km_\mathrm{g3p}} + \cfrac{[DXP]}{KM_\mathrm{dxp}} \right)}](/wiki/images/math/4/a/8/4a8faa589ce739bde2b441190759da1b.png)