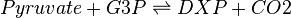Difference between revisions of "DXS"
Aliah.hawari (talk | contribs) (→DXS parameters) |
Aliah.hawari (talk | contribs) (→DXS parameters) |
||
| Line 35: | Line 35: | ||
| Kcat || Forward || DXS || 229.7 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || <ref name="Brammer2011"> [https://www.ncbi.nlm.nih.gov/pubmed/21878632 Brammer, L.A. 2011] "1-FDeoxy-D-xylulose 5-phosphate synthase catalyzes a novel random sequential mechanism", JBioChem, '''283'''(42):36522-36531.</ref> | | Kcat || Forward || DXS || 229.7 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || <ref name="Brammer2011"> [https://www.ncbi.nlm.nih.gov/pubmed/21878632 Brammer, L.A. 2011] "1-FDeoxy-D-xylulose 5-phosphate synthase catalyzes a novel random sequential mechanism", JBioChem, '''283'''(42):36522-36531.</ref> | ||
|- | |- | ||
| − | | Kcat || Forward || DXS || 153.6 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || Brammer2011 | + | | Kcat || Forward || DXS || 153.6 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || <ref>Brammer2011</ref> |
|- | |- | ||
| − | | Kcat || Forward || DXS || 145.5 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || Brammer2011 | + | | Kcat || Forward || DXS || 145.5 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min || <ref>Brammer2011</ref> |
|- | |- | ||
| − | | Kcat || Forward || DXS || 209 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || Brammer2011 | + | | Kcat || Forward || DXS || 209 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || <ref>Brammer2011</ref> |
|- | |- | ||
| − | | Kcat || Forward || DXS || 173 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || Brammer2011 | + | | Kcat || Forward || DXS || 173 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || <ref>Brammer2011</ref> |
|- | |- | ||
| − | | Kcat || Forward || DXS || 246 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || Brammer2011 | + | | Kcat || Forward || DXS || 246 || 1/min || 16 || from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min || <ref>Brammer2011</ref> |
|- | |- | ||
| Kcat || Forward || GAP || 48 || 1/min || 256 || Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. || Boronat1999 | | Kcat || Forward || GAP || 48 || 1/min || 256 || Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. || Boronat1999 | ||
Revision as of 13:51, 23 March 2017
You can go back to main page of the kinetic model here.
The DXS reaction (EC 2.2.1.7)
Deoxyxylulose-5-phosphate synthase (DXS) catalyses the production of 1-deoxy-D-xylulose 5-phosphate (DXP) from pyruvate and glyceraldehyde 3-phosphate (G3P). This reaction is the first step in the MEP pathway.
Modelling DXS
In the kinetic model, the DXS reaction is modelled with reversible Michaelis-Menten using the Hanekom [1] bi-bi random order generic equation. In total, this reaction requires five kinetic parameters (Kms for all substrates and products, and a forward Kcat) and one thermodynamic parameter (Equilibrium constant, Keq).
![V_\mathrm{DXS}= \cfrac{Kcat_\mathrm{forward} \bullet [DXS] \bullet \left( \cfrac{[Pyr]}{Km_\mathrm{DXS}} \right) \bullet \left( \cfrac{[G3P]}{Km_\mathrm{g3p}} \right) \bullet \left( 1 - \cfrac{\left( \cfrac{[DXP]\bullet[CO2]}{[Pyr]\bullet[G3P]} \right)}{K_\mathrm{eq}} \right)} {\left( 1 + \cfrac {[Pyr]}{Km_\mathrm{pyr}} + \cfrac{[CO2]}{Km_\mathrm{co2}}\right) \bullet \left( 1 + \cfrac{[G3P]}{Km_\mathrm{g3p}} + \cfrac{[DXP]}{KM_\mathrm{dxp}} \right)}](/wiki/images/math/4/a/8/4a8faa589ce739bde2b441190759da1b.png)
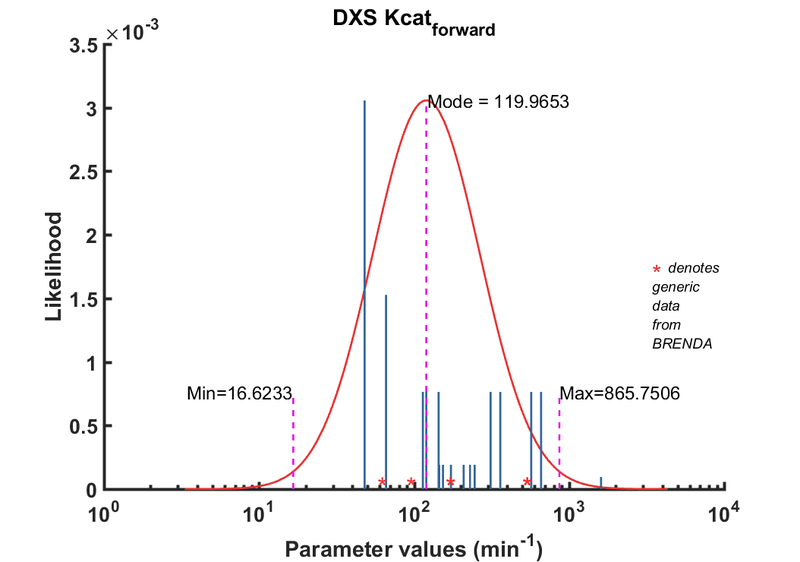
DXS parameters
| Parameter | Direction | Substrate | Value | Unit | Weight | Description | Reference |
|---|---|---|---|---|---|---|---|
| Kcat | Forward | DXS | 229.7 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [2] |
| Kcat | Forward | DXS | 153.6 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [3] |
| Kcat | Forward | DXS | 145.5 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 40mM Tris, pH 8, 37C ; Km_GAP:52.5 +/- 8.3 microM; Km_pyruvate: 86.3 +/- 16.2 microM, kcat: 145.50 +/- 12.7 1/min | [4] |
| Kcat | Forward | DXS | 209 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [5] |
| Kcat | Forward | DXS | 173 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [6] |
| Kcat | Forward | DXS | 246 | 1/min | 16 | from E. coli wild type DXS, with non-optimal buffer: 100mM Tris, pH 8, 37C ; Km_GAP:279.0+/- 7.2microM; Km_pyruvate: 74.70+/- 7.3 microM, kcat: 209.0 +/- 6.3 1/min | [7] |
| Kcat | Forward | GAP | 48 | 1/min | 256 | Taken from Cane 2001's ref20. E.coli DXS in 40mM Tris, pH7.5, 37¡C. Km pyruvate 2.9 ± 0.5 mM. | Boronat1999 |
| Kcat | Forward | GAP | 66 | 1/min | 128 | DXPS2; in vitro- S. coelicolor gene expressed in E. coli; pH 7.5, 47C. | Cane2001 |
| Kcat | Forward | GAP | 114 | 1/min | 8 | from R. capsulatus, pH 7.4, 37C | Eubanks2003, Br657953 |
| Kcat | Forward | GAP | 660 | 1/min | 64 | from Plasmodium, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | Handa2013, Br719510 |
| Kcat | Forward | GAP | 1608 | 1/min | 8 | dxs11 from Agrobacterium tumefaciens, pH8.0, 37¡C, expressed in E. coli | Lee2007, Br674982 |
| Kcat | Forward | GAP | 120 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Kcat | Forward | GAP | 120 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Kcat | Forward | GAP | 360 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Kcat | Forward | Pyruvate | 570 | 1/min | 64 | from Plasmodium, expressed in E. coli. Look at Table 3. pH7-7.5;37C, Km_GAp:19 +/- 4 microM; Km_Pyruvate: 870 +/- 110 microM. | Handa2013, Br719510 |
| Kcat | Forward | Pyruvate | 144 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Kcat | Forward | Pyruvate | 114 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
| Kcat | Forward | Pyruvate | 312 | 1/min | 64 | from Botrycoccus braunnii. Three recombinant enzymes used: DXS-I, DXS-II, DXS-III which are different by the digestion pattern using Xhol and BamHI. expressed in E. coli; pH 7.8 , 32 C; Km 1800 +/- 200 microM | Matsushima2012, Br720765 |
References
- ↑ Hanekom, A. J. 2006. "Generic kinetic equations for modelling multisubstrate reactions in computational systems biology", MSc Thesis submitted at the University of Stellenbosch
- ↑ Brammer, L.A. 2011 "1-FDeoxy-D-xylulose 5-phosphate synthase catalyzes a novel random sequential mechanism", JBioChem, 283(42):36522-36531.
- ↑ Brammer2011
- ↑ Brammer2011
- ↑ Brammer2011
- ↑ Brammer2011
- ↑ Brammer2011