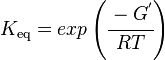Limonene Synthase
You can go back to main page of the kinetic model here.
Model construction progress :
| Enzyme | Pathway | Keq | Kcat | VmaxF | KmGPP | KmSub2 | [GPP] | [Sub2] | VmaxR | KmLim | KmPP | [Limonene] | [PP] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 02_LimSynth | Limonene Biosynthesis | 1 | 5 | 3 | 4 | 1 |
Legend:
| Have not started ·· | 1 -2 data found ·· | 3-4 data found ·· | sufficient data found/estimated ·· | data distribution generated ·· | data sampled |
| to do | DONE! |
Contents
What we know
Issues
Strategies
Reaction catalysed
Metabolite Background Information
Long metabolite names are abbreviated in the model for clarity and standard identification purposes.
| Metabolite | Abbreviation | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem |
|---|---|---|---|---|---|---|
| geranyl diphosphate | GPP | C10H20O7P2 | 314.209 | 17211 | 41432 | 445995 |
| (-)-4S-limonene | Limonene | C10H16 | 136.24 | 15384 | 449062 | 22311 or 439250 |
| diphosphate | PP | O7P2 | 173.94 | 644102 |
Equation Rate
| Parameter | Description | Units |
|---|---|---|
| VLimSynth | Reaction rate for Limonene Synthase | |
| Vmaxforward | Maximum reaction rate towards the production of limonene | ref |
| KmGPP | Michaelis-Menten constant for GPP | mM |
| KmLimonene | Michaelis-Menten constant for Limonene | mM |
| KmPP | Michaelis-Menten constant for PP | mM |
| Keq | Equilibrium constant | |
| [GPP] | GPP concentration | mM |
| [Limonene] | Limonene concentration | mM |
| [PP] | PP concentration | mM |
Strategies for estimating the kinetic parameter values
Calculating the Equilibrium Constant
Standard Gibbs Free energy
Standard Gibbs Free energy for Limonene Synthase from MetaCyc (EC 4.2.3.16) is -28.049988 kcal/mol [1].
SI derived unit for Gibbs free energy is Joules per mol (J mol-1). 1 kJ·mol−1 is equal to 0.239 kcal·mol−1.
Therefore, the Gibbs free energy for Limonene synthase in kJ mol-1 is:
- Failed to parse (Cannot store math image on filesystem.): \cfrac {1}{0.239 kcal.mol^-1} * -28.049988 kcal.mol^-1
- Failed to parse (Cannot store math image on filesystem.): = -117.36396 kJmol^-1
The Equilibrium Constant
The equilibrium constant can be calculated using the Van't Hoff Isotherm equation:
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac {-(- 117.36396 \text { kJmol}^{-1})}{ (8.31 \text{ JK}^{-1} \text { mol}^{-1} * 289 K} \right )
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac { + 117.36396 \text { kJmol}^{-1} }{ 2401.59 \text{ JK}^{-1}\text { mol}^{-1} }\right)
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac{ 117.364 * 10^3 \text { Jmol}^{-1}}{2401.59 \text{ JK}^{-1}\text { mol}^{-1}} \right)
Failed to parse (Cannot store math image on filesystem.): =exp \left ( 48.8693 \right )
Failed to parse (Cannot store math image on filesystem.): = 1.6736 * 10^{21}
where;
| Keq | Equilibrium constant |
| -ΔG° | Gibbs free energy change. For Limonene Synthase it is -117.364 kJmol-1 |
| R | Gas constant with a value of 8.31 JK-1mol-1 |
| T | Temperature which is always expressed in Kelvin |
Kinetic Parameter Values
Values for the kinetic parameter required to simulate this model can be obtained from published and unpublished literature.
A more detailed descriptions of the values listed above can be found here , where I've linked and highlighted where these data came from.
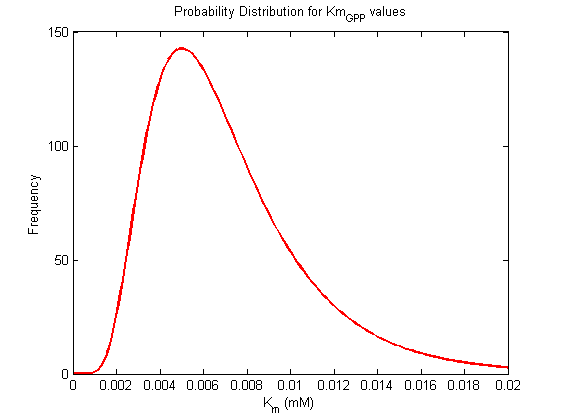
Km Values
| Km (mM) | Unit | Substrate / Product | Directionality | Organism | References |
|---|---|---|---|---|---|
| 0.00125 | mM | GPP | forward | Ricciocarpos natans | Ref |
| 0.0018 | mM | GPP | forward | Mentha piperita | Ref |
| 0.00625 | mM | GPP | forward | Cannabis sativa L. | Ref |
| 0.00496 | mM | GPP | forward | Cannabis sativa L. | Ref |
| 0.0031 | mM | GPP | forward | Citrus limon | ref |
| 0.016 | mM | GPP | forward | Escherichia coli | Ref |
| 0.0068 | mM | GPP | forward | Cannabis sativa L. | [2] |
| 0.0067 | mM | GPP | forward | Cannabis sativa L. | Ref |
Vmax values
| Vmax | Unit | Directionality | Organism | References |
|---|---|---|---|---|
| 0.08 | µmol/min/mg | forward | Cannabis sativa L. | References |
| 0.13 | µmol/min/mg | forward | Cannabis sativa L. | References |
| 0.4748 | µmol/min/mg | forward | Citrus limon | References |
| Vmax | Unit | Directionality | Organism | References |
| Vmax | Unit | Directionality | Organism | References |
| Vmax | Unit | Directionality | Organism | References |
Kcat values
| Kcat | Unit | Organism | Reference |
|---|---|---|---|
| 0.3 | s-1 | Mentha piperita & Mentha spicata | [3] |
| 0.08 | s-1 | Cannabis sativa L. | Reference |
| 0.14 | s-1 | Cannabis sativa L. | Reference |
| 0.02 | s-1 | E. coli | Reference |
| 0.082 | s-1 | Cannabis sativa L. | [2] |
Extracting Information from Limonene Production Rates
The production rates would reflect on the flux for this enzyme, and this would provide provide the insights on the Vmax of this enzyme.
| Amount produced (mg/L) | Time (H) | Organism | Description | Reaction Flux (µM/s) |
|---|---|---|---|---|
| 5 | 24 | Escherichia coli | Possible reason for the low limonene production might due to the insufficient supply of IPP and DMAPP [4]. | 0.0255 |
| 335 | 48 | Escherichia coli | Engineered E.coli in which heterologous MVA pathway was installed [5]. | 0.8537 |
| 35.8 | 48 | Escherichia coli | E.coli was engineered to express GPPS, LS, DXS, and IDI [6] . | 0.0912 |
| 4.87 | 48 | Escherichia coli | This was the initial titer. The study established a limonene biosynthesis pathway in E.coli using four different polycistronic operons based on 3 vectors with varied expression strength [6]. | 0.0124 |
| 17.4 | 48 | Escherichia coli | Using a plasmid with DXS and IDI over expressed [6]. | 0.0445 |
| 430 | 72 | Escherichia coli | [5] | 0.7306 |
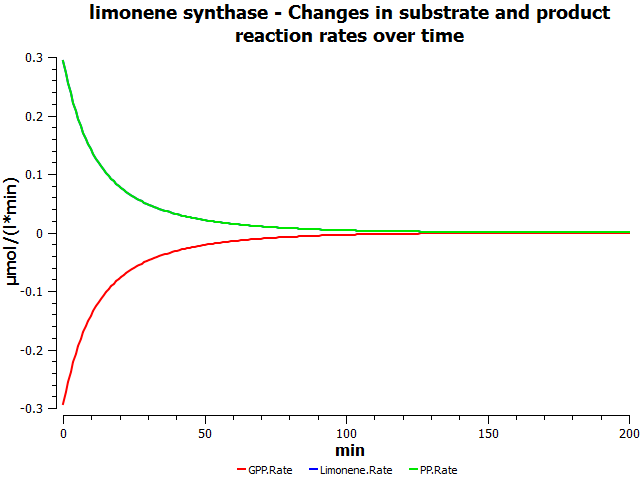
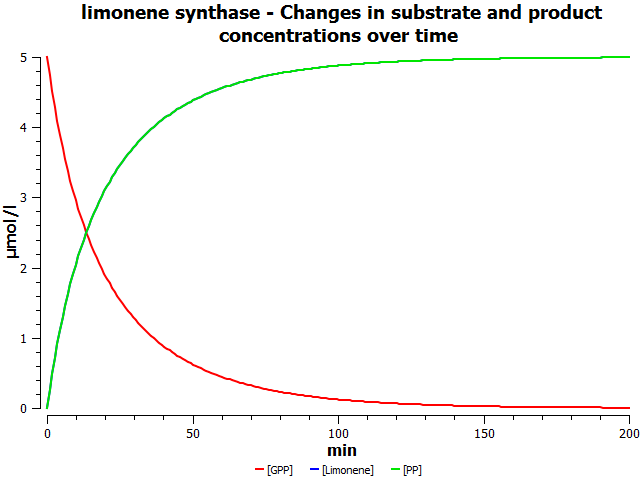
Simulations
these are mock simulation results
References
- ↑ Latendresse M. (2013). "Computing Gibbs Free Energy of Compounds and Reactions in MetaCyc."
- ↑ 2.0 2.1 Günnewich et. al. 2006. "Functional expression and characterization of trichome-specific (-)-limonene synthase and (+)-α-pinene synthase from Cannabis sativa", Natural Product Communications, 0(0):1-10
- ↑ Alonso et. al. 1992. "Purification of 4S-Limonene Synthase, a Monoterpene Cyclase from the Glandular Trichomes of Peppermint (Mentha x piperita) and Spearmint (Mentha spicata)", The Journal of Biological Chemistry, 267(11):7582-7587
- ↑ Carter, Ora A. et. al.2013. "Monoterpene biosynthesis pathway construction in Escherichia coli",Phytochemistry, 64:425–433, 2003.
- ↑ 5.0 5.1 Alonso-Gutierez et. al. 2013. "Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production", Metabolic Engineering, 19:33-41 Cite error: Invalid
<ref>tag; name "AlonsoGutierez2013" defined multiple times with different content - ↑ 6.0 6.1 6.2 Du et. al. 2014. "Enhanced limonene production by optimizing the expression of limonene biosynthesis and MEP pathway genes in E.coli", Bioprocessing and Bioprocessing, 1:10
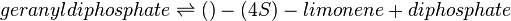
![V_\mathrm{LimSynth} = Vmax_\mathrm{forward} * \cfrac {\cfrac{[GPP]}{Km_\mathrm{GPP}} * \left ( 1 - \cfrac {[Limonene]*[PP]}{[GPP]*K_\mathrm{eq}} \right )}{1 + \cfrac {[GPP]}{Km_\mathrm{GPP}} + \cfrac {[Limonene]}{Km_\mathrm{Limonene}} + \cfrac {[PP]}{Km_\mathrm{PP}} + \cfrac {[Limonene]*[PP]}{Km_\mathrm{Limonene}*Km_\mathrm{PP}}}](/wiki/images/math/6/9/8/698f16353b522504a844aae5c37bdac9.png)