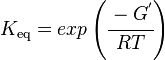Limonene Synthase
You can go back to main page of the kinetic model here.
Contents
What we know
Issues
Strategies
Reaction catalysed
Metabolite Background Information
Long metabolite names are abbreviated in the model for clarity and standard identification purposes.
| Metabolite | Abbreviation | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem |
|---|---|---|---|---|---|---|
| geranyl diphosphate | GPP | C10H20O7P2 | 314.209 | 17211 | 41432 | 445995 |
| (-)-4S-limonene | Limonene | C10H16 | 136.24 | 15384 | 449062 | 22311 or 439250 |
| diphosphate | PP | O7P2 | 173.94 | 644102 |
Equation Rate
| Parameter | Description | Reference |
|---|---|---|
| VLimSynth | Reaction rate for Limonene Synthase | ref |
| Vmaxforward | Maximum reaction rate towards the production of limonene | ref |
| KmGPP | Michaelis-Menten constant for GPP | ref |
| KmLimonene | Michaelis-Menten constant for Limonene | ref |
| KmPP | Michaelis-Menten constant for PP | ref |
| Keq | Equilibrium constant | ref |
| [GPP] | GPP concentration | ref |
| [Limonene] | Limonene concentration | ref |
| [PP] | PP concentration | ref |
Strategies for estimating the kinetic parameter values
Calculating the Equilibrium Constant
The equilibrium constant can be calculated using the Van't Hoff Isotherm equation:
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac {-(- 117.36396 \text { kJmol}^{-1})}{ (8.31 \text{ JK}^{-1} \text { mol}^{-1} * 289 K} \right )
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac { + 117.36396 \text { kJmol}^{-1} }{ 2401.59 \text{ JK}^{-1}\text { mol}^{-1} }\right)
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac{ 117.364 * 10^3 \text { Jmol}^{-1}}{2401.59 \text{ JK}^{-1}\text { mol}^{-1}} \right)
Failed to parse (Cannot store math image on filesystem.): =exp \left ( 48.8693 \right )
Failed to parse (Cannot store math image on filesystem.): = 1.6736 * 10^{21}
where;
| Keq | Equilibrium constant |
| -ΔG° | Gibbs free energy change. For Limonene Synthase it is -117.364 kJmol-1 |
| R | Gas constant with a value of 8.31 JK-1mol-1 |
| T | Temperature which is always expressed in kelvin |
Standard Gibbs Free energy
Standard Gibbs Free energy for Limonene Synthase from MetaCyc (EC 4.2.3.16) is -28.049988 kcal/mol [1].
SI derived unit for Gibbs free energy is Joules per mol (J mol-1). 1 kJ·mol−1 is equal to 0.239 kcal·mol−1.
Therefore, the Gibbs free energy for Limonene synthase in kJ mol-1 is:
- Failed to parse (Cannot store math image on filesystem.): \cfrac {1}{0.239 kcal.mol^-1} * -28.049988 kcal.mol^-1
- Failed to parse (Cannot store math image on filesystem.): = -117.36396 kJmol^-1
Extracting Information from Limonene Production Rates
Production rates would reflect the flux for this reaction in the forward direction.
| Amount produced (mg/L) | Time (H) | Organism | Description | Reaction Flux (µM/s) |
|---|---|---|---|---|
| 5 | 24 | Escherichia coli | Possible reason for the low limonene production might due to the insufficient supply of IPP and DMAPP [2]. | 0.0255 |
| 335 | 48 | Escherichia coli | Engineered E.coli in which heterologous MVA pathway was installed [3]. | 0.8537 |
| 35.8 | 48 | Escherichia coli | E.coli was engineered to express GPPS, LS, DXS, and IDI [4] . | 0.0912 |
| 4.87 | 48 | Escherichia coli | This was the initial titer. The study established a limonene biosynthesis pathway in E.coli using four different polycistronic operons based on 3 vectors with varied expression strength [5]. | 0.0124 |
| 17.4 | 48 | Escherichia coli | Using a plasmid with DXS and IDI over expressed [6]. | 0.0445 |
| 430 | 72 | Escherichia coli | [7] | 0.7306 |
Published Kinetic Parameter Values
| Km (mM) | Vmax | Kcat (s-1) | Kcat/Km | Organism | Description |
|---|---|---|---|---|---|
| 0.00125 | - | - | - | Ricciocarpos natans | GPP -> Limonene |
| 0.0018 | - | - | - | Mentha piperita | GPP -> Limonene |
| 0.00625 | 0.08 µmol/min/mg | 0.08 | 1.5 | Cannabis sativa L. | GPP -> Limonene |
| 0.00496 | 0.13 µmol/min/mg | 0.14 | 2.9 | Cannabis sativa L. | GPP -> Limonene |
| 0.0031 | 0.4748 µmol/min/mg | - | - | Citrus limon | GPP -> Limonene |
| 0.016 | - | 0.02 | - | Escherichia coli | GPP -> Limonene |
| 0.0068 | - | 0.082 | - | Cannabis sativa L. | GPP -> Limonene |
| 0.0067 | - | 0.081 | - | Cannabis sativa L. | GPP -> Limonene |
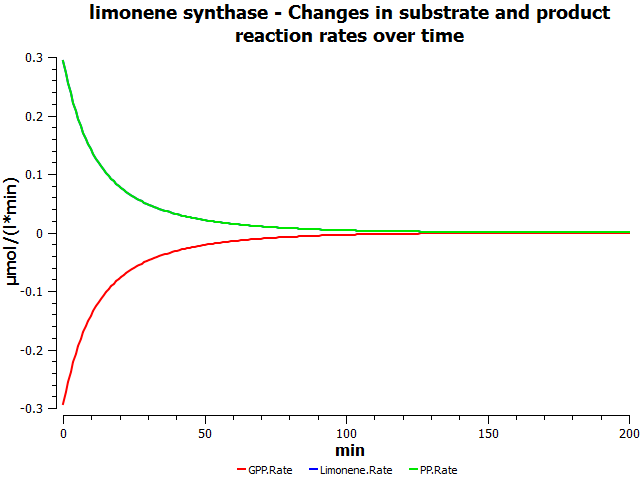
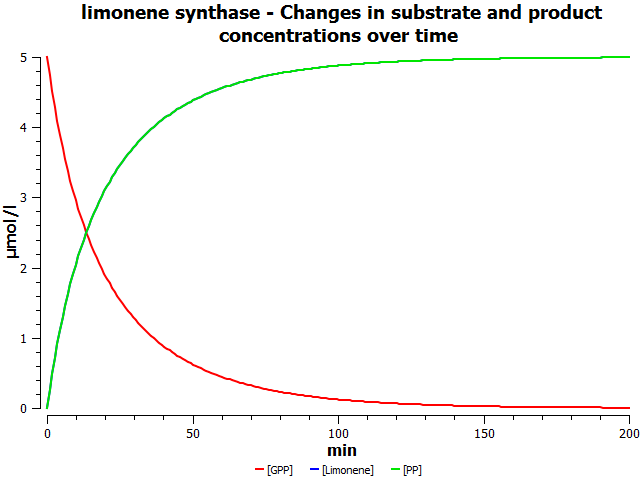
Simulations
References
- ↑ Latendresse M. (2013). "Computing Gibbs Free Energy of Compounds and Reactions in MetaCyc."
- ↑ Carter, Ora A. et. al.2013. "Monoterpene biosynthesis pathway construction in Escherichia coli",Phytochemistry, 64:425–433, 2003.
- ↑ Alonso-Gutierez et. al. 2013. "Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production", Metabolic Engineering, 19:33-41
- ↑ Du et. al. 2014. "Enhanced limonene production by optimizing the expression of limonene biosynthesis and MEP pathway genes in E.coli", Bioprocessing and Bioprocessing, 1:10
- ↑ Du et. al. 2014. "Enhanced limonene production by optimizing the expression of limonene biosynthesis and MEP pathway genes in E.coli", Bioprocessing and Bioprocessing, 1:10
- ↑ Du et. al. 2014. "Enhanced limonene production by optimizing the expression of limonene biosynthesis and MEP pathway genes in E.coli", Bioprocessing and Bioprocessing, 1:10
- ↑ Alonso-Gutierez et. al. 2013. "Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production", Metabolic Engineering, 19:33-41
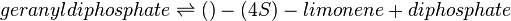
![V_\mathrm{LimSynth} = Vmax_\mathrm{forward} * \cfrac {\cfrac{[GPP]}{Km_\mathrm{GPP}} * \left ( 1 - \cfrac {[Limonene]*[PP]}{[GPP]*K_\mathrm{eq}} \right )}{1 + \cfrac {[GPP]}{Km_\mathrm{GPP}} + \cfrac {[Limonene]}{Km_\mathrm{Limonene}} + \cfrac {[PP]}{Km_\mathrm{PP}} + \cfrac {[Limonene]*[PP]}{Km_\mathrm{Limonene}*Km_\mathrm{PP}}}](/wiki/images/math/6/9/8/698f16353b522504a844aae5c37bdac9.png)