Mitocondrial pyruvate metabolism
Mitocondrial pyruvate metabolism(MPM) is an enzyme that generates ATP form pyruvate.
Contents
Chemical reaction
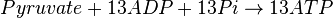
Rate equation
- Chemical reactions proceed to equilibrium within closed systems. For a simple reaction
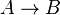 it is defined as
it is defined as ![K_{eq} = \frac{[B]_{eq}}{[A]_{eq}}](/wiki/images/math/a/c/8/ac844b70f8f7c5f1d91accb00061a1ea.png) where forward and reverse rates are equal.
where forward and reverse rates are equal. - Equilibrium is not reached in open system due to influx and outflux. Mass action ratio[1]
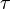 for
for 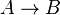 reaction is defined as
reaction is defined as ![\tau = \frac{[B]_{ob}}{[A]_{ob}}](/wiki/images/math/8/0/5/8050a810298db95df036d87c2e79f1ea.png) where subscript ob represents observable at a given point.
where subscript ob represents observable at a given point. - Deviation from equilibrium is measured with Disequilibrium constant
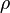 as
as 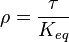
- Given the simple uni molecular reaction
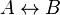 the mass action equation can be modified as
the mass action equation can be modified as
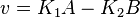
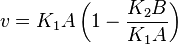
Considering 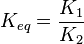 and
and 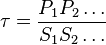 we have,
we have,
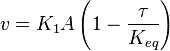
- The generalized arbitrary mass action ratio gives us
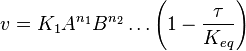
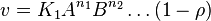
For eg. for the reaction 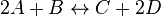 , the rate law would be
, the rate law would be 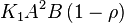
- This equation demonstrates how a rate expression can be divided into parts that include both kinetics and thermodynamic properties [2].
- Given the net rate of reaction
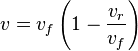 , we have
, we have
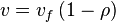
In this model
- The rate law is defined as
![v = K_1[Pyruvate][ADP]^{13}[Pi]^{13}\left(1-\frac{\frac{[ATP]^{13}}{[Pyruvate][ADP]^{13}[Pi]^{13}}}{K_{eq}}\right)](/wiki/images/math/c/0/7/c07d97f8c92fac816401bb1210a69046.png)
- The overall standard free-energy change for Pyruvate metabolism is Failed to parse (Cannot store math image on filesystem.): \Delta G^o{'}= -30.5 Kj/Mol [3].
- Calculating
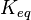 value from these free energy gives Failed to parse (Cannot store math image on filesystem.): \Delta G' = - 30.5\ kJ.mol^{-1}
, Failed to parse (Cannot store math image on filesystem.): Keq = exp(-\frac{\Delta G'}{RT}) = exp(\frac{30500}{8.31*298.15}) \approx 221941.39
value from these free energy gives Failed to parse (Cannot store math image on filesystem.): \Delta G' = - 30.5\ kJ.mol^{-1}
, Failed to parse (Cannot store math image on filesystem.): Keq = exp(-\frac{\Delta G'}{RT}) = exp(\frac{30500}{8.31*298.15}) \approx 221941.39
- Calculating
- The Flux of pyruvate consumed by mitochondria measured for AS_30D is
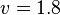 [4].
[4]. - The steady state concentrations for substrates and products are
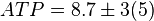 ,
, 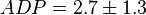 ,
, 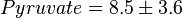 and
and 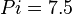 .
. - The
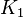 value calculated from the above mentioned values are Failed to parse (Cannot store math image on filesystem.): 2.20E-018
value calculated from the above mentioned values are Failed to parse (Cannot store math image on filesystem.): 2.20E-018
- To calculate the uncertainty of
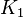 we first looked at the uncertainty on the substrate and product concentration. The maximum uncertainty reported for these values are
we first looked at the uncertainty on the substrate and product concentration. The maximum uncertainty reported for these values are 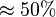 . In our model we considered this
. In our model we considered this 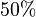 uncertainty in its mean value giving value of Failed to parse (Cannot store math image on filesystem.): 2.20E^{-018} \pm 1.099E^{-018}
uncertainty in its mean value giving value of Failed to parse (Cannot store math image on filesystem.): 2.20E^{-018} \pm 1.099E^{-018}
Parameter values
| Parameter | Value | Organism | Remarks |
|---|---|---|---|
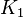
|
Failed to parse (Cannot store math image on filesystem.): 2.20E^{-018} |
Parameters with uncertainty
| Parameter | Value | Organism | Remarks |
|---|---|---|---|
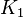
|
Failed to parse (Cannot store math image on filesystem.): 2.20E^{-018} \pm 1.099E^{-018} |
References
- ↑ Hess B. and Brand K. (1965), Enzymes and metabolite profiles. In Control of energy metabolism. III. Ed. B. Chance, R. K. Estabrook and J. R. Williamson. New York: Academic Press
- ↑ Sauro H M, Enzyme Kinetics for Systems Biology, Second Edition, Ambrosius Publishing (2013), ISBN-10: 0-9824773-3-3
- ↑ Nelson D. and Cox M. (2008), Lehninger Principles of Biochemistry, Fight Edition, W.H. Freeman and Company, ISBN-10: 071677108X
- ↑ Marín-Hernández A, Gallardo-Pérez JC, Rodríguez-Enríquez S et al (2011) Modeling cancer glycolysis. Biochim Biophys Acta 1807:755–767 (doi)