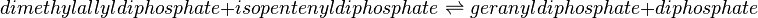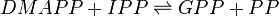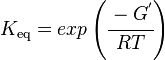Difference between revisions of "Geranyl diphosphate synthase (GPPS)"
Aliah.hawari (talk | contribs) (→Strategies for estimating the kinetic parameter values) |
Aliah.hawari (talk | contribs) (→Calculating the Equilibrium Constant) |
||
| Line 155: | Line 155: | ||
<math> | <math> | ||
| − | = exp \left ( \cfrac {-( | + | = exp \left ( \cfrac {-(-13.0 \text { kcalmol}^{-1})}{ (1.98722 \text{ calK}^{-1} \text { mol}^{-1} * 289 K} \right ) |
</math> | </math> | ||
| Line 161: | Line 161: | ||
<math> | <math> | ||
| − | = exp \left ( \cfrac { | + | = exp \left ( \cfrac {13.0 \text { kcalmol}^{-1} }{ 574.30658 \text{ calK}^{-1}\text { mol}^{-1} }\right) |
</math> | </math> | ||
<math> | <math> | ||
| − | = exp \left ( \cfrac{ | + | = exp \left ( \cfrac{ 13000 \text {calmol}^{-1}}{574.30658 \text{ JK}^{-1}\text { mol}^{-1}} \right) |
</math> | </math> | ||
| Line 192: | Line 192: | ||
| T || Temperature which is always expressed in kelvin, as a standard, I used 298 K | | T || Temperature which is always expressed in kelvin, as a standard, I used 298 K | ||
|} | |} | ||
| − | |||
| − | |||
| − | |||
=== Published Kinetic Parameter Values === | === Published Kinetic Parameter Values === | ||
Revision as of 13:13, 24 March 2016
You can go back to main page of the kinetic model here.
Legend:
| Have not started ·· | 1 -2 data found ·· | 3-4 data found ·· | sufficient data found/estimated ·· | data distribution generated ·· | data sampled |
| to do | DONE! |
Contents
What we know
Issues
Strategies
Reaction catalysed
Metabolite and Enzyme Background Information
Long metabolite names are abbreviated in the model for clarity and standard identification purposes.
| Metabolite | Abbreviation | Chemical Formula | Molar mass (g/mol) | ChEBI | ChEMBL | PubChem | MetaCyc |
|---|---|---|---|---|---|---|---|
| dimethylallyl diphosphate | DMAPP | ||||||
| isopentenyl diphosphate | IPP | ||||||
| geranyl diphosphate | GPP | C10H20O7P2 | 314.209 | 17211 | 41432 | 445995 | |
| diphosphate | PP | O7P2 | 173.94 | 644102 | |||
| geranyl diphosphate synthase | GPPS | 32.16 kD (from nucleotide sequence), 36 kD (experimental) | CPLX-8656 |
Equation Rate
- Failed to parse (Cannot store math image on filesystem.): V_\mathrm{GPPS} = Vmax_\mathrm{forward} * \cfrac { \left (\cfrac{[DMAPP]}{Km_\mathrm{DMAPP}} * \cfrac{[IPP]}{Km_\mathrm{IPP}}\right )* \left ( 1 - \cfrac {[GPP]*[PP]}{[GPP]*[PP]*K_\mathrm{eq}} \right )}{\left (1 + \cfrac {[IPP]}{Km_\mathrm{IPP}} + \cfrac {[PP]}{Km_\mathrm{PP}} \right ) * \left ( 1 + \cfrac {[DMAPP]}{Km_\mathrm{DMAPP}} + \cfrac {[GPP]}{Km_\mathrm{GPP}} \right )}
| Parameter | Description |
|---|---|
| VGPPS | Reaction rate for Geranyl diphosphate synthase |
| Vmaxforward | Maximum reaction rate towards the production of GPP |
| KmGPP | Michaelis-Menten constant for GPP |
| KmIPP | Michaelis-Menten constant for IPP |
| KmPP | Michaelis-Menten constant for PP |
| KmDMAPP | Michaelis-Menten constant for DMAPP |
| Keq | Equilibrium constant |
| [GPP] | GPP concentration |
| [DMAPP] | DMAPP concentration |
| [IPP] | DMAPP concentration |
| [PP] | PP concentration |
Strategies for estimating the kinetic parameter values
Calculating the Equilibrium Constant
Standard Gibbs Free energy for GPPS from MetaCyc is -13.0 kcal mol-1.
The equilibrium constant can be calculated using the Van't Hoff Isotherm equation:
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac {-(-13.0 \text { kcalmol}^{-1})}{ (1.98722 \text{ calK}^{-1} \text { mol}^{-1} * 289 K} \right )
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac {13.0 \text { kcalmol}^{-1} }{ 574.30658 \text{ calK}^{-1}\text { mol}^{-1} }\right)
Failed to parse (Cannot store math image on filesystem.): = exp \left ( \cfrac{ 13000 \text {calmol}^{-1}}{574.30658 \text{ JK}^{-1}\text { mol}^{-1}} \right)
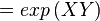
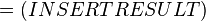
where;
| Keq | Equilibrium constant |
| -?G° | Gibbs free energy change. For GPPS it is -13.0 kcalmol-1 |
| R | Gas constant with a value of 1.98722 calK-1mol-1 |
| T | Temperature which is always expressed in kelvin, as a standard, I used 298 K |
Published Kinetic Parameter Values
Km Values
| Km (mM) | Unit | Substrate / Product | Directionality | Organism | References |
|---|---|---|---|---|---|
| Value | unit | substrate | directionality | organism | Ref |
Vmax values
| Vmax | Unit | Directionality | Organism | References |
|---|---|---|---|---|
| Value | µmol/min/mg (unit) | directionality | Organism | References |
Kcat values
| Kcat | Unit | Organism | Reference |
|---|---|---|---|
| value | s-1 | Organism | ref e.g. Alonso 1992 [1] |
Extracting Information from (INSERT SUBSTRATE/PRODUCT) Production Rates
| Amount produced (mg/L) | Time (H) | Organism | Description | Reaction Flux (µM/s) |
|---|---|---|---|---|
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |
| X | X | Y | Z | Z |