Difference between revisions of "Phosphoserine amino-transferase"
(→Parameters with uncertainty) |
(→Parameters with uncertainty) |
||
| Line 41: | Line 41: | ||
* serC is the enzyme concentration collected from Turnaev ''et. al.'' <ref name="Turnaev_2006"></ref> which is considered to be fixed. | * serC is the enzyme concentration collected from Turnaev ''et. al.'' <ref name="Turnaev_2006"></ref> which is considered to be fixed. | ||
* Mean and standard deviation for <math>Kc_{PHP}</math> and <math>Kc_{PSER}</math> is reported in <ref name="Ali_2006">Ali, V., and T. Nozaki, (2006), ''Biochemical and functional characterization of phosphoserine aminotransferase from Entamoeba histolytica, which possesses both phosphorylated and non-phosphorylated serine metabolic pathways'', Mol. Biochem. Parasitol. 145:71-83</ref> for ''Entamoeba histolytica''(an anaerobic parasitic protozoan). The values are <math>Kc_{PHP} = 0.058 \pm 0.021 </math> and <math>Kc_{PSER} = 0.0368 \pm 0.004 </math> | * Mean and standard deviation for <math>Kc_{PHP}</math> and <math>Kc_{PSER}</math> is reported in <ref name="Ali_2006">Ali, V., and T. Nozaki, (2006), ''Biochemical and functional characterization of phosphoserine aminotransferase from Entamoeba histolytica, which possesses both phosphorylated and non-phosphorylated serine metabolic pathways'', Mol. Biochem. Parasitol. 145:71-83</ref> for ''Entamoeba histolytica''(an anaerobic parasitic protozoan). The values are <math>Kc_{PHP} = 0.058 \pm 0.021 </math> and <math>Kc_{PSER} = 0.0368 \pm 0.004 </math> | ||
| + | *For the turnover number (<math>KcatC</math>)10% Std. Dev. is considered. | ||
| + | |||
{|class="wikitable" | {|class="wikitable" | ||
! Parameter | ! Parameter | ||
| Line 55: | Line 57: | ||
|- | |- | ||
|<math>KcatC</math> | |<math>KcatC</math> | ||
| − | |1.75 | + | |<math>1.75 \pm 0.175 </math> |
|1/s | |1/s | ||
|- | |- | ||
Revision as of 17:18, 6 May 2014
This enzyme catalyzes reaction to convert Phosphohydroxypyruvate (PHP) to phosphoserine (PSER).
Contents
Chemical equation
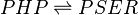
Rate equation
Special rate equation [1] is used.
![\frac{serC * KcatC * \frac{[PHP]}{Kc_{PHP}} }{ 1 + \frac{[PHP]}{Kc_{PHP}} + \frac{[PSER]}{Kc_{PSER}} }](/wiki/images/math/8/c/c/8ccb0a98d76ad1a351009f4c75ae7a7f.png)
Parameter values
| Parameter | Value | Units | Organism | Remarks |
|---|---|---|---|---|
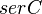
|
0.1 [2] | mM | Escherichia coli | |
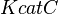
|
1.75 [3] | 1/s | ||
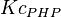
|
0.0015[3] | mM | ||
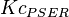
|
0.0017[3] | mM |
Parameters with uncertainty
- serC is the enzyme concentration collected from Turnaev et. al. [2] which is considered to be fixed.
- Mean and standard deviation for
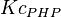 and
and 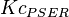 is reported in [4] for Entamoeba histolytica(an anaerobic parasitic protozoan). The values are
is reported in [4] for Entamoeba histolytica(an anaerobic parasitic protozoan). The values are 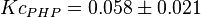 and
and 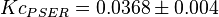
- For the turnover number (
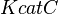 )10% Std. Dev. is considered.
)10% Std. Dev. is considered.
| Parameter | Value | Units | Organism | Remarks |
|---|---|---|---|---|
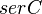
|
0.1 [2] | mM | Escherichia coli | |
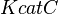
|

|
1/s | ||
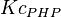
|
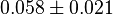
|
mM | Entamoeba histolytica | |
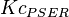
|
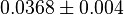
|
mM | Entamoeba histolytica |
References
- ↑ Smallbone K, Stanford NJ (2013). Kinetic modeling of metabolic pathways: Application to serine biosynthesis. In: Systems Metabolic Engineering, Humana Press. pp. 113–121
- ↑ 2.0 2.1 2.2 Turnaev II, Ibragimova SS, Usuda Y et al (2006). Mathematical modeling of serine and glycine synthesis regulation in Escherichia coli. Proceedings of the fifth international conference on bioinformatics of genome regulation and structure 2:78–83
- ↑ 3.0 3.1 3.2 Drewke C, Klein M, Clade D et al (1996) 4-O-phosphoryl-L-threonine, a substrate of the pdxC(serC) gene product involved in vitamin B6 biosynthesis. FEBS Lett 390:179–182
- ↑ Ali, V., and T. Nozaki, (2006), Biochemical and functional characterization of phosphoserine aminotransferase from Entamoeba histolytica, which possesses both phosphorylated and non-phosphorylated serine metabolic pathways, Mol. Biochem. Parasitol. 145:71-83