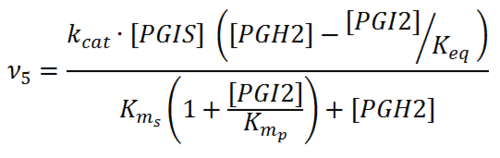Difference between revisions of "Transformation of PGH2 to PGI2"
(→Parameters) |
(→Parameters) |
||
| (19 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
[[Welcome to the In-Silico Model of Cutaneous Lipids Wiki | Return to overview]] | [[Welcome to the In-Silico Model of Cutaneous Lipids Wiki | Return to overview]] | ||
| + | |||
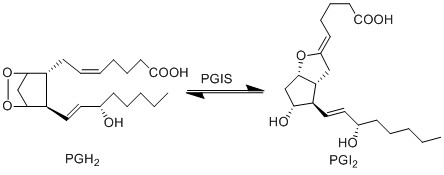
| + | The isomerisation of PGH2 to prostacyclin (PGI2), is performed by prostaglandin I synthase (PGIS). This protein is a member of the CYP P450 family, but unlike most CYP P450 enzymes it does not oxidise PGH2. PGI2 is generated by the rearrangement of the peroxide functional group, whereby a hydroxyl group is formed at C11, and a new epoxide ring is formed between C9 and C6. PGI2 is generated by the sequential action of COX and PGIS, which co-localise in the ER, plasma membrane and nuclear membrane <ref>Smith, W. L. DeWitt, D. L. Allen, M. L. , ''Bimodal distribution of the prostaglandin I2 synthase antigen in smooth muscle cells'', J Biol Chem (1983), 258, 5922-6.</ref>. | ||
| + | |||
== Reaction == | == Reaction == | ||
[[File:R5_PGH2_-_PGI2.jpg|center|500px]] | [[File:R5_PGH2_-_PGI2.jpg|center|500px]] | ||
| Line 5: | Line 8: | ||
==Chemical equation== | ==Chemical equation== | ||
| − | <center><math> | + | <center><math> PGH2 \rightleftharpoons PGI2 </math></center> |
== Rate equation == | == Rate equation == | ||
| + | [[File:R05.PNG|center|500px]] | ||
== Parameters == | == Parameters == | ||
| + | === K<sub>ms</sub> === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 19: | Line 24: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 24: | Line 30: | ||
|<math> mM </math> | |<math> mM </math> | ||
|Human | |Human | ||
| − | |Human PGIS | + | |Expression Vector: Bovine Endothelial and Aorta Cells |
| + | Enzyme: Human PGIS | ||
| + | pH:7.4 | ||
| + | Temperature: 23 | ||
| + | |256 | ||
|<ref name="Yeh2005"> [http://www.ncbi.nlm.nih.gov/pubmed/16406803 H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase.'' Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.]</ref> | |<ref name="Yeh2005"> [http://www.ncbi.nlm.nih.gov/pubmed/16406803 H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase.'' Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.]</ref> | ||
|- | |- | ||
| Line 30: | Line 40: | ||
|<math> mM </math> | |<math> mM </math> | ||
|Bovine | |Bovine | ||
| − | |Bovine Endothelial Cells | + | |Expression Vector: Bovine Endothelial and Aorta Cells |
| − | |<ref name=" | + | Enzyme: Bovine PGIS |
| + | pH:7.4 | ||
| + | Temperature: 24 | ||
| + | |192 | ||
| + | |<ref name="Hara19941"> [http://www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGIS Kms distribution | ||
| + | ! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.24E-02 || 2.46E+00 || -4.22E+00 || 4.20E-01 | ||
| + | |} | ||
| + | |||
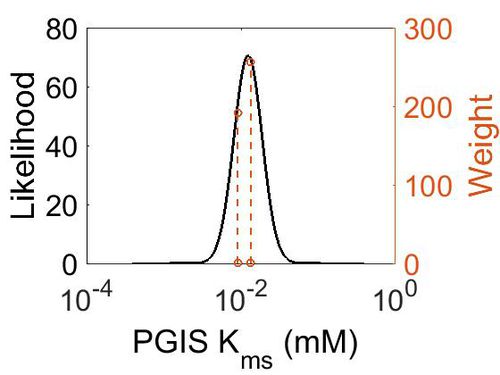
| + | [[Image:17.jpg|none|thumb|500px|The estimated probability distribution for PGIS Kms. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === K<sub>mp</sub> === | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGIS Kmp distribution | ||
| + | ! Mode (mM) !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.29E-02 || -4.182022605 || 0.414183521 | ||
| + | |} | ||
| + | |||
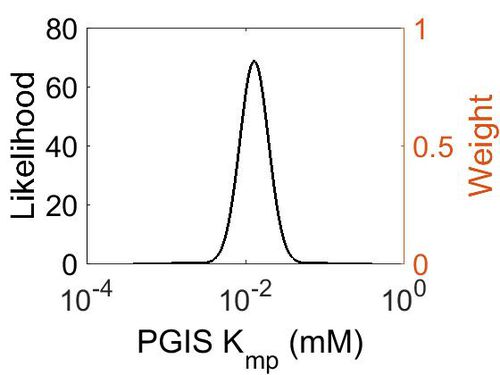
| + | [[Image:18.jpg|none|thumb|500px|The estimated probability distribution for PGIS Kmp. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | |||
| + | ===k<sub>cat</sub> === | ||
{|class="wikitable sortable" | {|class="wikitable sortable" | ||
| − | |+ style="text-align: left;" | | + | |+ style="text-align: left;" | Literature values |
|- | |- | ||
! Value | ! Value | ||
| Line 42: | Line 79: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 47: | Line 85: | ||
| per minute | | per minute | ||
| Cattle | | Cattle | ||
| − | | Bovine | + | |Expression Vector: E. Coli |
| − | | <ref name=" | + | Enzyme: Bovine PGIS |
| + | pH:7.4 | ||
| + | Temperature: 24 | ||
| + | |192 | ||
| + | | <ref name="Hara19942"> [http://www.ncbi.nlm.nih.gov/pubmed/8051072 Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells.'' J Biol Chem. 1994 Aug 5;269(31):19897-903.]</ref> | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGIS kcat distribution | ||
| + | ! Mode (min-1) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.41E+02 || 1.35E+00 || 5.028213628 || 0.287348692 | ||
|} | |} | ||
| + | |||
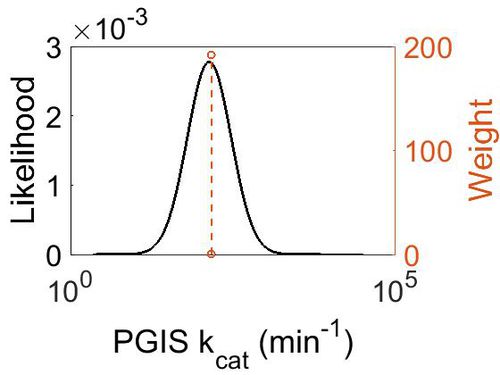
| + | [[Image:19.jpg|none|thumb|500px|The estimated probability distribution for PGIS kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === Enzyme concentration === | ||
| + | |||
| + | To convert the enzyme concentration from ppm to mM, the following [[Common equations#Enzyme concentration (mM)|equation]] was used. | ||
| + | |||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | |- | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | |412 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Urinary bladder | ||
| + | Enzyme: PGIS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
| + | |- | ||
| + | |206 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Lung | ||
| + | Enzyme: PGIS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
| + | |- | ||
| + | |60.1 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Esophagus | ||
| + | Enzyme: PGIS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
| + | |- | ||
| + | |9.93 | ||
| + | |<math> ppm </math> | ||
| + | |Human | ||
| + | |Expression Vector: Oral Cavity | ||
| + | Enzyme: PGIS | ||
| + | pH: 7.5 | ||
| + | Temperature: 37 °C | ||
| + | |1024 | ||
| + | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGIS concentration distribution | ||
| + | ! Mode (ppm) !! Mode (mM) !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 1.13E+02 ||6.25E-04 || 4.13E+00 || 5.67E+00 || 9.68E-01 | ||
| + | |- | ||
| + | |} | ||
| + | |||
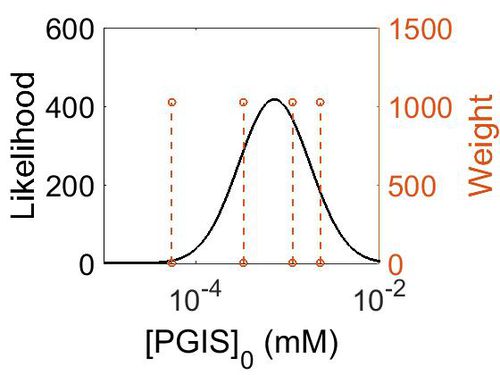
| + | [[Image:164.jpg|none|thumb|500px|The estimated probability distribution for PGIS concentration. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
| + | |||
| + | === K<sub>eq</sub> === | ||
| + | {|class="wikitable sortable" | ||
| + | |+ style="text-align: left;" | Literature values | ||
| + | |- | ||
| + | ! Gibbs Free Energy Change | ||
| + | ! Units | ||
| + | ! Species | ||
| + | ! Notes | ||
| + | ! Weight | ||
| + | ! Reference | ||
| + | |- | ||
| + | | 0.04 | ||
| + | |kcal/mol | ||
| + | |Not stated | ||
| + | |Estimated | ||
| + | Enzyme: Transacylase | ||
| + | Substrate: | ||
| + | Product: | ||
| + | pH: 7.3 | ||
| + | ionic strength: 0.25 | ||
| + | |64 | ||
| + | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=PROSTAGLANDIN-I-SYNTHASE-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |+ style="text-align: left;" | Description of the PGIS Keq distribution | ||
| + | ! Mode !! Confidence Interval !! Location parameter (µ) !! Scale parameter (σ) | ||
| + | |- | ||
| + | | 9.35E-01 || 1.00E+01 || 7.30E-01 || 8.90E-01 | ||
| + | |} | ||
| + | |||
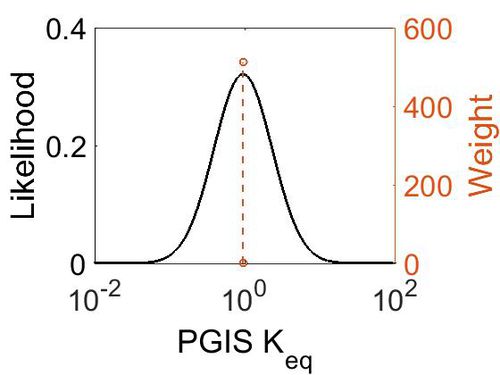
| + | [[Image:20.jpg|none|thumb|500px|The estimated probability distribution for PGIS kcat. The value and weight of the literature values used to define the distribution are indicated by an orange dashed line. The x axis is plotted on a log-scale. ]] | ||
== References == | == References == | ||
Latest revision as of 09:58, 2 November 2019
The isomerisation of PGH2 to prostacyclin (PGI2), is performed by prostaglandin I synthase (PGIS). This protein is a member of the CYP P450 family, but unlike most CYP P450 enzymes it does not oxidise PGH2. PGI2 is generated by the rearrangement of the peroxide functional group, whereby a hydroxyl group is formed at C11, and a new epoxide ring is formed between C9 and C6. PGI2 is generated by the sequential action of COX and PGIS, which co-localise in the ER, plasma membrane and nuclear membrane [1].
Contents
Reaction
Chemical equation
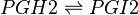
Rate equation
Parameters
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 1.33E-02 ± 1.40E-03 | 
|
Human | Expression Vector: Bovine Endothelial and Aorta Cells
Enzyme: Human PGIS pH:7.4 Temperature: 23 |
256 | [2] |
| 9.00E-03 ± 5.00E-03 | 
|
Bovine | Expression Vector: Bovine Endothelial and Aorta Cells
Enzyme: Bovine PGIS pH:7.4 Temperature: 24 |
192 | [3] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.24E-02 | 2.46E+00 | -4.22E+00 | 4.20E-01 |
Kmp
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 1.29E-02 | -4.182022605 | 0.414183521 |
kcat
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 147 ± 45 | per minute | Cattle | Expression Vector: E. Coli
Enzyme: Bovine PGIS pH:7.4 Temperature: 24 |
192 | [4] |
| Mode (min-1) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 1.41E+02 | 1.35E+00 | 5.028213628 | 0.287348692 |
Enzyme concentration
To convert the enzyme concentration from ppm to mM, the following equation was used.
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 412 | 
|
Human | Expression Vector: Urinary bladder
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| 206 | 
|
Human | Expression Vector: Lung
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| 60.1 | 
|
Human | Expression Vector: Esophagus
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| 9.93 | 
|
Human | Expression Vector: Oral Cavity
Enzyme: PGIS pH: 7.5 Temperature: 37 °C |
1024 | [6] |
| Mode (ppm) | Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|---|
| 1.13E+02 | 6.25E-04 | 4.13E+00 | 5.67E+00 | 9.68E-01 |
Keq
| Gibbs Free Energy Change | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 0.04 | kcal/mol | Not stated | Estimated
Enzyme: Transacylase Substrate: Product: pH: 7.3 ionic strength: 0.25 |
64 | [7] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 9.35E-01 | 1.00E+01 | 7.30E-01 | 8.90E-01 |
References
- ↑ Smith, W. L. DeWitt, D. L. Allen, M. L. , Bimodal distribution of the prostaglandin I2 synthase antigen in smooth muscle cells, J Biol Chem (1983), 258, 5922-6.
- ↑ H. C. Yeh, P. Y. Hsu, Characterization of heme environment and mechanism of peroxide bond cleavage in human prostacyclin synthase. Biochim Biophys Acta. 2005 Dec 30;1738(1-3):121-32.
- ↑ Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells. J Biol Chem. 1994 Aug 5;269(31):19897-903.
- ↑ Hara S., Isolation and molecular cloning of prostacyclin synthase from bovine endothelial cells. J Biol Chem. 1994 Aug 5;269(31):19897-903.
- ↑ 5.0 5.1 5.2 M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471