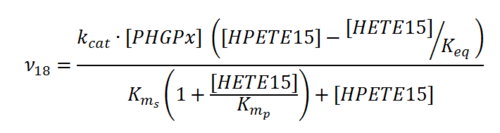Difference between revisions of "Transformation of 15-HPETE to 15-HETE"
(→Parameters) |
(→Parameters) |
||
| Line 26: | Line 26: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 35: | Line 36: | ||
pH: 7 | pH: 7 | ||
Temperature: 37 | Temperature: 37 | ||
| + | |512 | ||
|<ref name="Liu2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18163571 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein.'' Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.]</ref> | |<ref name="Liu2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18163571 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein.'' Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.]</ref> | ||
|- | |- | ||
| Line 44: | Line 46: | ||
pH: 7 | pH: 7 | ||
Temperature: 37 | Temperature: 37 | ||
| + | |1024 | ||
|<ref name="Zheng2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18373941 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases.'' Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.]</ref> | |<ref name="Zheng2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18373941 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases.'' Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.]</ref> | ||
|- | |- | ||
| Line 53: | Line 56: | ||
pH: 7.4 | pH: 7.4 | ||
Temperature: 37 | Temperature: 37 | ||
| − | + | |768 | |
|<ref name="Hirat1997"> [http://www.ncbi.nlm.nih.gov/pubmed/9030530 Hiratsuka A. "Subunit Ya-specific glutathione peroxidase activity toward cholesterol 7-hydroperoxides of glutathione S-transferases in cytosols from rat liver and skin.'' J Biol Chem. 1997 Feb 21;272(8):4763-9.]</ref> | |<ref name="Hirat1997"> [http://www.ncbi.nlm.nih.gov/pubmed/9030530 Hiratsuka A. "Subunit Ya-specific glutathione peroxidase activity toward cholesterol 7-hydroperoxides of glutathione S-transferases in cytosols from rat liver and skin.'' J Biol Chem. 1997 Feb 21;272(8):4763-9.]</ref> | ||
|- | |- | ||
| Line 87: | Line 90: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 96: | Line 100: | ||
pH: 7 | pH: 7 | ||
Temperature: 37 | Temperature: 37 | ||
| + | |512 | ||
|<ref name="Liu2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18163571 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein.'' Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.]</ref> | |<ref name="Liu2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18163571 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein.'' Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.]</ref> | ||
|- | |- | ||
| Line 105: | Line 110: | ||
pH: 7 | pH: 7 | ||
Temperature: 37 | Temperature: 37 | ||
| + | |1024 | ||
|<ref name="Zheng2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18373941 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases.'' Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.]</ref> | |<ref name="Zheng2008"> [http://www.ncbi.nlm.nih.gov/pubmed/18373941 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases.'' Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.]</ref> | ||
|- | |- | ||
| Line 111: | Line 117: | ||
{| class="wikitable" | {| class="wikitable" | ||
|+ style="text-align: left;" | Description of the PHGPx kcat distribution | |+ style="text-align: left;" | Description of the PHGPx kcat distribution | ||
| − | ! Mode (min-1) !! Confidence Interval !! | + | ! Mode (min-1) !! Confidence Interval !! Locati1024on parameter (µ) !! Scale parameter (σ) |
|- | |- | ||
| 2.44E+04 || 4.20E+00 || 1.11E+01 || 9.75E-01 | | 2.44E+04 || 4.20E+00 || 1.11E+01 || 9.75E-01 | ||
| Line 127: | Line 133: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 136: | Line 143: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |2048 | ||
|<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
|- | |- | ||
| Line 145: | Line 153: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
|- | |- | ||
| Line 154: | Line 163: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | |<ref name="Wilhelm2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13319.pdf M. Wilhelm ''Mass-spectrometry-based draft of the human proteome'' Nature, 2014 509, 582–587]</ref> | ||
|- | |- | ||
| Line 163: | Line 173: | ||
pH: 7.5 | pH: 7.5 | ||
Temperature: 37 °C | Temperature: 37 °C | ||
| + | |1024 | ||
|<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | |<ref name="Kim2014"> [http://www.nature.com/nature/journal/v509/n7502/pdf/nature13302.pdf M. Kim ''A draft map of the human proteome'' Nature, 2014 509, 575–581]</ref> | ||
|- | |- | ||
| Line 185: | Line 196: | ||
! Species | ! Species | ||
! Notes | ! Notes | ||
| + | ! Weight | ||
! Reference | ! Reference | ||
|- | |- | ||
| Line 196: | Line 208: | ||
pH: 7.3 | pH: 7.3 | ||
ionic strength: 0.25 | ionic strength: 0.25 | ||
| + | |64 | ||
|<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=1.11.1.12-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | |<ref name="MetaCyc”>[http://metacyc.org/META/NEW-IMAGE?type=REACTION&object=1.11.1.12-RXN Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471]</ref> | ||
|} | |} | ||
Revision as of 10:51, 22 May 2019
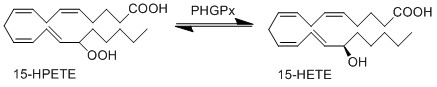
15-HPETE is reduced by phospholipid hydroperoxide glutathione peroxidase (PHGPx) to the corresponding alcohol, 15-HETE (Suth01). 15-HETE and 12-HETE have reciprocal activity during inflammation, for example 15-HETE reduces the infiltration of inflammatory cells, meanwhile 12-HETE acts as an activator for infiltrating immune cells (Dowd, Kobza Black et al. 1985, Serhan, Jain et al. 2003).
Contents
Reaction
Chemical equation
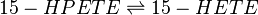
Rate equation
Parameters
Note that the literature values are the same as reaction 12.
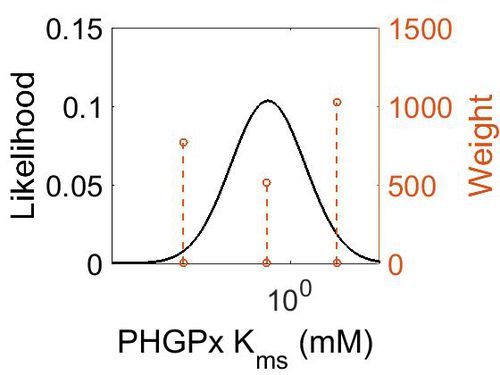
Kms
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 3.00E-01 | 
|
Human | Expression Vector: Bioimprited Enzyme - Selenosubtilisin
Enzyme: Glutathione Peroxidase pH: 7 Temperature: 37 |
512 | [1] |
| 11.1 ± 2.90E-01 | 
|
Human | Expression Vector: E Coli
Enzyme: Wild Type Glutathione Peroxidase Enzyme (Se-hGSTZ1-1) pH: 7 Temperature: 37 |
1024 | [2] |
| 4.00E-03 | 
|
Rat | Expression Vector: Rat Liver Cells
Enzyme: Glutathione Peroxidase pH: 7.4 Temperature: 37 |
768 | [3] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 3.19E-01 | 1.02E+03 | 2.54E+00 | 1.92E+00 |
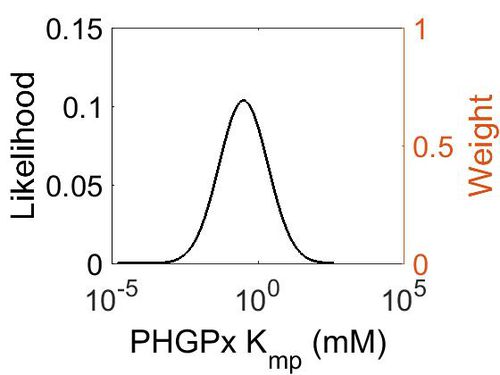
Kmp
This is a “Dependent parameter”, meaning that the log-normal distribution for this parameter was calculated using multivariate distributions (this is discussed in detail here). As a result, no confidence interval factor or literature values were cited for this parameter.
| Mode (mM) | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|
| 3.15E-01 | 2.53E+00 | 1.92E+00 |
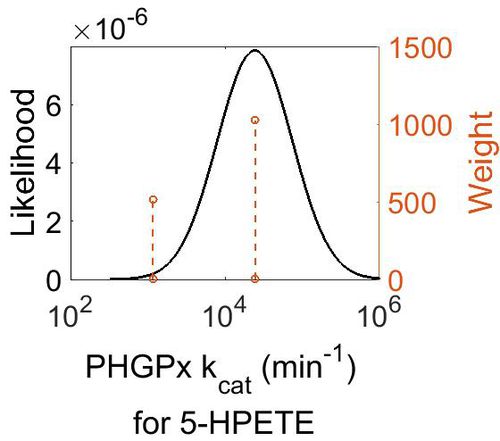
kcat
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 1170 ± 50 | 
|
Human | Expression Vector: Bioimprited Enzyme - Selenosubtilisin
Enzyme: Glutathione Peroxidase pH: 7 Temperature: 37 |
512 | [1] |
| 24500 ± 150 | 
|
Human | Expression Vector: E Coli
Enzyme: Wild Type Glutathione Peroxidase Enzyme (Se-hGSTZ1-1) pH: 7 Temperature: 37 |
1024 | [2] |
| Mode (min-1) | Confidence Interval | Locati1024on parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 2.44E+04 | 4.20E+00 | 1.11E+01 | 9.75E-01 |
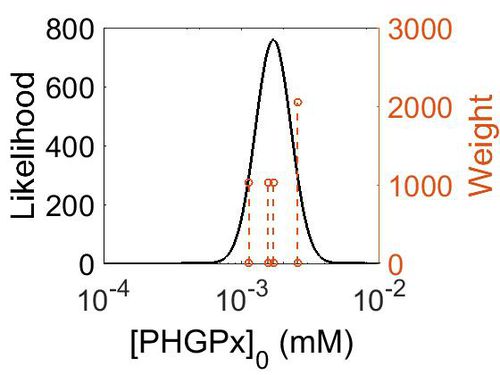
Enzyme concentration
| Value | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| 459 | 
|
Human | Expression Vector: Skin
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
2048 | [4] |
| 307 | 
|
Human | Expression Vector: Lung
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| 282 | 
|
Human | Expression Vector: Esophagus
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
1024 | [4] |
| 204 | 
|
Human | Expression Vector: Gut
Enzyme: PHGPx pH: 7.5 Temperature: 37 °C |
1024 | [5] |
| Mode (mM) | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 3.14E+02 | 1.38E+00 | 5.85E+00 | 3.09E-01 |
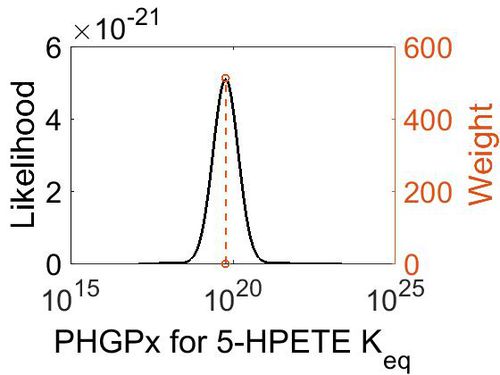
Keq
| Gibbs Free Energy Change | Units | Species | Notes | Weight | Reference |
|---|---|---|---|---|---|
| (-26.941177) | kcal/mol | Not stated | Estimated
Enzyme: PHGPx Substrate:a hydroperoxy-fatty-acyl-[lipid] Product: a hydroxy-fatty-acyl-[lipid] pH: 7.3 ionic strength: 0.25 |
64 | [6] |
| Mode | Confidence Interval | Location parameter (µ) | Scale parameter (σ) |
|---|---|---|---|
| 5.90E+19 | 1.00E+01 | 4.63E+01 | 8.90E-01 |
Related Reactions
- ↑ 1.0 1.1 Liu L. "Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein. Biomacromolecules. 2008 Jan;9(1):363-8. doi: 10.1021/bm7008312. Epub 2007 Dec 29.
- ↑ 2.0 2.1 Zheng K. "A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases. Int J Biochem Cell Biol. 2008;40(10):2090-7. doi: 10.1016/j.biocel.2008.02.006. Epub 2008 Feb 15.
- ↑ Hiratsuka A. "Subunit Ya-specific glutathione peroxidase activity toward cholesterol 7-hydroperoxides of glutathione S-transferases in cytosols from rat liver and skin. J Biol Chem. 1997 Feb 21;272(8):4763-9.
- ↑ 4.0 4.1 4.2 M. Wilhelm Mass-spectrometry-based draft of the human proteome Nature, 2014 509, 582–587
- ↑ M. Kim A draft map of the human proteome Nature, 2014 509, 575–581
- ↑ Caspi et al 2014, "The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases," Nucleic Acids Research 42:D459-D471