Difference between revisions of "Nucleosid diphosphate kinase"
| (21 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
| + | [[Category:Uncertainty]] | ||
'''Nucleoside-diphosphate kinase'''s are enzymes that catalyze the exchange of phosphate groups between different nucleotides. The overall effect of NDKs is to transfer a phosphate group from a nucleoside triphosphate to a nucleoside diphosphate. Starting with ATP and UDP, the activity of NDK produces ADP and UTP. | '''Nucleoside-diphosphate kinase'''s are enzymes that catalyze the exchange of phosphate groups between different nucleotides. The overall effect of NDKs is to transfer a phosphate group from a nucleoside triphosphate to a nucleoside diphosphate. Starting with ATP and UDP, the activity of NDK produces ADP and UTP. | ||
| Line 5: | Line 6: | ||
==Rate equation== | ==Rate equation== | ||
| − | Random order Bi-Bi rate law is used | + | Random order Bi-Bi rate law is used from <ref name="Konig_2012">M. König, S. Bulik, H.G. Holzhütter (2012), ''Quantifying the contribution of the liver to glucose homeostasis: a detailed kinetic model of human hepatic glucose metabolism'', PLoS Comput. Biol., 8 (6), p. e1002577</ref> |
| − | <center><math>\frac{ \frac{V_{max}}{K_m^{ATP}K_m^{UDP}}\left( [ATP][UDP] - \frac{[ADP][UTP]}{K_{eq}} \right) } {1}</math></center> | + | <center><math>\frac{ \frac{V_{max}}{K_m^{ATP}K_m^{UDP}}\left( [ATP][UDP] - \frac{[ADP][UTP]}{K_{eq}} \right) } { \left( 1 + \frac{[ATP]}{K_{m}^{ATP}} \right)\left( 1 + \frac{[UDP]}{K_{m}^{UDP}} \right) + \left( 1 + \frac{[ADP]}{K_{m}^{ADP}} \right)\left( 1 + \frac{[UTP]}{K_{m}^{UTP}} \right) -1 }</math></center> |
| + | |||
| + | |||
| + | ==Parameter values== | ||
| + | * A mean body weight is defined as 75Kg in Konig et. al <ref name="Konig_2012"></ref>. The <math>V_{max}</math> is calculated by multiplying the reported value with body weight. | ||
| + | |||
| + | {|class="wikitable" | ||
| + | ! Parameter | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Organism | ||
| + | ! Remarks | ||
| + | |- | ||
| + | |<math>V_{max}</math> | ||
| + | |<math>131.25</math> <ref name = "villar_1960"> Villar-Palasi C & Larner J (1960). ''Levels of activity of the enzymes of the glycogen cycle in rat tissues''. Arch Biochem Biophys 86, 270–273.</ref> | ||
| + | |<math>min^{-1}</math> | ||
| + | |Human Liver | ||
| + | |rowspan="6"| | ||
| + | |- | ||
| + | |<math>K_{m}^{ATP}</math> | ||
| + | |<math> 1.33 </math><ref name="Walter_1980">Walter P and Blobel G. (1980), ''Purification of a membrane-associated protein complex required for protein translocation across the endoplasmic reticulum'', Proc. Natl. Acad. Sci. 77:7112-7116</ref> | ||
| + | |mM | ||
| + | |Rat liver | ||
| + | |- | ||
| + | |<math>K_{m}^{ADP}</math> | ||
| + | |<math>0.042</math><ref name="Walter_1980"></ref> | ||
| + | |mM | ||
| + | |Rat liver | ||
| + | |- | ||
| + | |<math>K_{m}^{UTP}</math> | ||
| + | |<math>16</math><ref name="Fukuchi_1994"> Fukuchi, T., Shimada, N., Hanai, N., Ishikawa, N., Watanabe, K., and Kimura, N. (1994) ''Recombinant rat nucleoside diphosphate kinase isoforms (<math>\alpha</math> and <math>\beta</math>): purification, properties and application to immunological detection of native isoforms in rat tissues. Biochim. Biophys. Acta 1205, 113–122</ref> | ||
| + | |mM | ||
| + | |Rat tissue | ||
| + | |- | ||
| + | |<math>K_{m}^{UDP}</math> | ||
| + | |<math>0.19</math><ref name="Walter_1980"></ref> | ||
| + | |mM | ||
| + | |Rat liver | ||
| + | |- | ||
| + | |<math>K_{eq}</math> | ||
| + | |<math>1</math> | ||
| + | |Dimensionless | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | ==Parameters with uncertainty== | ||
| + | * In the paper by Walter (1980) et. al. <ref name="Walter_1980"></ref> two values of <math>K_{m}^{ADP}</math>, <math>K_{m}^{ATP}</math> and <math>K_{m}^{UDP}</math> have been reported; one for cytosolic kinase and the other for membrane-associated kinase. As in our model we are not differentiating between cytosolic and membrane-associated kinase, we would consider both those values and calculate the uncertainty from them. | ||
| + | |||
| + | * Similarly two values of <math>K_{m}^{UTP}</math> has been reported in Fukuchi (1994) et. al. <ref name="Fukuchi_1994"></ref>; for <math>\alpha</math> kinase and <math>\beta</math> kinase. Taking average of those kinase gives a value of <math>21.5 \pm 7.77</math> | ||
| + | |||
| + | * Two values of <math>V_{max}</math> is also reported in Fukuchi (1994) et. al. <ref name="Fukuchi_1994"></ref>; 126 (4.20 <math>\text{mmol min}^{-1}\text{ mg}^{-1}</math> multiplied with 30 mg) and 138 (4.60 <math>\text{mmol min}^{-1}\text{ mg}^{-1}</math> multiplied with 30 mg) <math>\text{mmol min}^{-1}</math>. Taking average gives <math>132 \pm 8.48</math>. <br>'''Note:''' This <math>V_{max}</math> value fails to reach a Steady-State for our model. Therefore we collected the <math>V_{max}</math> for ''Saccharomyces cerevisiae'' from Jong(1991) et. al. <ref name="Jong_1990"> Ambrose Y. Jong, Jin. J. Ma (1991), ''Saccharomyces cerevisiae nucleoside-diphosphate kinase: Purification, characterization, and substrate specificity'', Archives of Biochemistry and Biophysiscs, Vol. 291, No. 2, pp. 241-246</ref> which is <math>8.5 \times 10^{-6}</math>. But the value does not report any uncertainty. We took the largest Relative percente error of <math>V_{max}</math> in the model. Which is <math>71%</math>. Therefore the value with uncertainty becomes <math> 8.5 \times 10^{-6} \pm 6.04 \times 10^{-6} </math> | ||
| + | |||
| + | *Standard free energy <math>\Delta G^{o}{'}</math> is reported to be zero kJ/mol. Therefore the value of Keq can be assumed to be constant. As there was no other value reported, Std. Dev. from transporter Keq values are considered which is <math>\approx 70%</math>. So the Keq value would be <math>1 \pm 0.7</math> | ||
| + | |||
| + | {|class="wikitable" | ||
| + | ! Parameter | ||
| + | ! Value | ||
| + | ! Units | ||
| + | ! Organism | ||
| + | ! Remarks | ||
| + | |- | ||
| + | |<math>V_{max}</math> | ||
| + | |<math>8.5 \times 10^{-6} \pm 6.04 \times 10^{-6}</math> | ||
| + | |<math>min^{-1}</math> | ||
| + | |rowspan="6"|Rat lever | ||
| + | |rowspan="6"| | ||
| + | |- | ||
| + | |<math>K_{m}^{ATP}</math> | ||
| + | |<math> 1.49 \pm 0.23 </math> | ||
| + | |mM | ||
| + | |- | ||
| + | |<math>K_{m}^{ADP}</math> | ||
| + | |<math>0.0575 \pm 0.02</math> | ||
| + | |mM | ||
| + | |- | ||
| + | |<math>K_{m}^{UTP}</math> | ||
| + | |<math>21.5 \pm 7.77</math> | ||
| + | |mM | ||
| + | |- | ||
| + | |<math>K_{m}^{UDP}</math> | ||
| + | |<math>0.17 \pm 0.02</math> | ||
| + | |mM | ||
| + | |- | ||
| + | |<math>K_{eq}</math> | ||
| + | |<math>1 \pm 0.7</math> | ||
| + | |Dimensionless | ||
| + | |} | ||
| + | |||
| + | ==References== | ||
| + | <references/> | ||
Latest revision as of 11:38, 17 July 2014
Nucleoside-diphosphate kinases are enzymes that catalyze the exchange of phosphate groups between different nucleotides. The overall effect of NDKs is to transfer a phosphate group from a nucleoside triphosphate to a nucleoside diphosphate. Starting with ATP and UDP, the activity of NDK produces ADP and UTP.
Contents
Chemical equation
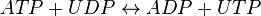
Rate equation
Random order Bi-Bi rate law is used from [1]
![\frac{ \frac{V_{max}}{K_m^{ATP}K_m^{UDP}}\left( [ATP][UDP] - \frac{[ADP][UTP]}{K_{eq}} \right) } { \left( 1 + \frac{[ATP]}{K_{m}^{ATP}} \right)\left( 1 + \frac{[UDP]}{K_{m}^{UDP}} \right) + \left( 1 + \frac{[ADP]}{K_{m}^{ADP}} \right)\left( 1 + \frac{[UTP]}{K_{m}^{UTP}} \right) -1 }](/wiki/images/math/9/e/6/9e6a0b14129bad329a3c09d3c7a69f43.png)
Parameter values
- A mean body weight is defined as 75Kg in Konig et. al [1]. The
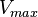 is calculated by multiplying the reported value with body weight.
is calculated by multiplying the reported value with body weight.
| Parameter | Value | Units | Organism | Remarks |
|---|---|---|---|---|
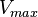
|
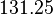 [2] [2]
|

|
Human Liver | |
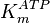
|
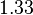 [3] [3]
|
mM | Rat liver | |
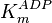
|
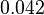 [3] [3]
|
mM | Rat liver | |
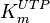
|
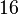 [4] [4]
|
mM | Rat tissue | |
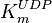
|
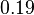 [3] [3]
|
mM | Rat liver | |
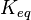
|
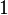
|
Dimensionless |
Parameters with uncertainty
- In the paper by Walter (1980) et. al. [3] two values of
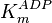 ,
, 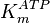 and
and 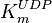 have been reported; one for cytosolic kinase and the other for membrane-associated kinase. As in our model we are not differentiating between cytosolic and membrane-associated kinase, we would consider both those values and calculate the uncertainty from them.
have been reported; one for cytosolic kinase and the other for membrane-associated kinase. As in our model we are not differentiating between cytosolic and membrane-associated kinase, we would consider both those values and calculate the uncertainty from them.
- Similarly two values of
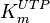 has been reported in Fukuchi (1994) et. al. [4]; for
has been reported in Fukuchi (1994) et. al. [4]; for 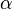 kinase and
kinase and 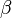 kinase. Taking average of those kinase gives a value of
kinase. Taking average of those kinase gives a value of 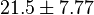
- Two values of
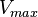 is also reported in Fukuchi (1994) et. al. [4]; 126 (4.20
is also reported in Fukuchi (1994) et. al. [4]; 126 (4.20 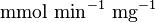 multiplied with 30 mg) and 138 (4.60
multiplied with 30 mg) and 138 (4.60 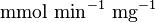 multiplied with 30 mg)
multiplied with 30 mg) 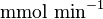 . Taking average gives
. Taking average gives 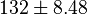 .
.
Note: This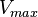 value fails to reach a Steady-State for our model. Therefore we collected the
value fails to reach a Steady-State for our model. Therefore we collected the 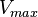 for Saccharomyces cerevisiae from Jong(1991) et. al. [5] which is
for Saccharomyces cerevisiae from Jong(1991) et. al. [5] which is 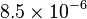 . But the value does not report any uncertainty. We took the largest Relative percente error of
. But the value does not report any uncertainty. We took the largest Relative percente error of 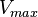 in the model. Which is
in the model. Which is 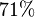 . Therefore the value with uncertainty becomes
. Therefore the value with uncertainty becomes 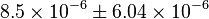
- Standard free energy
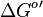 is reported to be zero kJ/mol. Therefore the value of Keq can be assumed to be constant. As there was no other value reported, Std. Dev. from transporter Keq values are considered which is
is reported to be zero kJ/mol. Therefore the value of Keq can be assumed to be constant. As there was no other value reported, Std. Dev. from transporter Keq values are considered which is 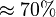 . So the Keq value would be
. So the Keq value would be 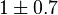
| Parameter | Value | Units | Organism | Remarks |
|---|---|---|---|---|
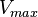
|
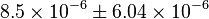
|

|
Rat lever | |
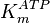
|
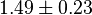
|
mM | ||
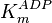
|
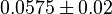
|
mM | ||
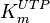
|
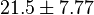
|
mM | ||
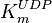
|
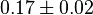
|
mM | ||
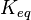
|
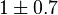
|
Dimensionless |
References
- ↑ 1.0 1.1 M. König, S. Bulik, H.G. Holzhütter (2012), Quantifying the contribution of the liver to glucose homeostasis: a detailed kinetic model of human hepatic glucose metabolism, PLoS Comput. Biol., 8 (6), p. e1002577
- ↑ Villar-Palasi C & Larner J (1960). Levels of activity of the enzymes of the glycogen cycle in rat tissues. Arch Biochem Biophys 86, 270–273.
- ↑ 3.0 3.1 3.2 3.3 Walter P and Blobel G. (1980), Purification of a membrane-associated protein complex required for protein translocation across the endoplasmic reticulum, Proc. Natl. Acad. Sci. 77:7112-7116
- ↑ 4.0 4.1 4.2 Fukuchi, T., Shimada, N., Hanai, N., Ishikawa, N., Watanabe, K., and Kimura, N. (1994) Recombinant rat nucleoside diphosphate kinase isoforms (
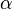 and
and 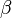 ): purification, properties and application to immunological detection of native isoforms in rat tissues. Biochim. Biophys. Acta 1205, 113–122
): purification, properties and application to immunological detection of native isoforms in rat tissues. Biochim. Biophys. Acta 1205, 113–122
- ↑ Ambrose Y. Jong, Jin. J. Ma (1991), Saccharomyces cerevisiae nucleoside-diphosphate kinase: Purification, characterization, and substrate specificity, Archives of Biochemistry and Biophysiscs, Vol. 291, No. 2, pp. 241-246