Welcome to the In-Silico Model of butyrolactone regulation in Streptomyces coelicolor
Contents
About the Project
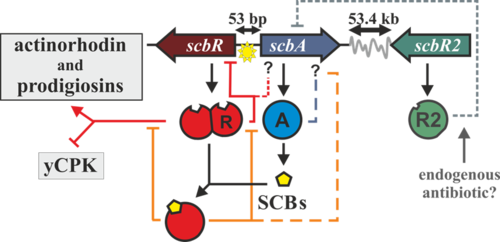
Streptomyces are Gram-positive soil-dwelling bacteria, which are known for being prolific sources of secondary metabolites, many of which have medical interest (e.g.: streptomycin, cloramphenicol or kanamycin). Recent genome sequencing, and posterior bioinformatic analysis, using tools such as antiSMASH [[1]], have shown the presence of putative cryptic secondary metabolite-producing clusters. One way Streptomyces can coordinate secondary metabolite production is through the use of small diffusible molecules, known as γ-butyrolactones. In the model organism Streptomyces coelicolor the butyrolactone regulatory system involves a synthase (scbA) and a butyrolactone receptor (scbR).
The scbA gene is presumed to catalyse the condensation of dihydroxyacetone phosphate with a beta-ketoacid to produce three different butyrolactones (SCB1, SCB2 and SCB3. Hereafter named as SCBs). On the other hand, scbR is a TetR-like DNA-binding protein known to regulate its own transcription and that of scbA and to directly regulate production of a cryptic metabolite and indirectly regulate production of blue pigmented actinorhodin (Act) and prodigiosins (e.g. undecylprodigiosin, Red). These genes are divergently encoded and their promoter regions overlap 53bp.
Transcription analysis have shown that both genes are mainly active during transition from logaritmic growth to stationary phase. It is presumed that SCBs slowly accumulate into the media and upon reaching a concentration threshold promote a coordinated switch-like transition to antibiotic production by binding to ScbR. However, the mechanism of this network is not fully defined, although several alternative scenarios have been proposed. In 2008, Mehra et al. [1] proposed a deterministic model involving a putative ScbR-ScbA complex. More recently, Chatterjee et al. [2] proposed that the promoter overlap between scbA and scbR is the sole driving force of the precise switch-like transition. More detailed information on the GBL system and the previous modelling work can be found in the section Background Information on GBL system.
The aim of this project is the design and analysis of a stochastic and parameter uncertainty-aware model which describes the butyrolactone regulatory system and allows reliable predictions of its behaviour. The complete elucidation of this system could potentially lead to the design of robust and sensitive systems as orthologous regulatory circuits in synthetic biology and biotechnology.
Description of the Model
Several alternative scenarios for the mechanism of action of the GBL system have been previously proposed. Our aim is to create a unified model which will include them all and enable their parallel or combined analysis. The scenarios investigated are the following:
- TI – The effect of transcriptional interference (collisions between the elongating RNAPs which leads to transcriptional termination) due to the overlap of the two genes’ promoter regions by 53 bp and by the convergent transcription of the two genes. This results in a decrease in expression of full-length mRNAs from both promoters and production of truncated mRNAs. Note: The transcriptional interference mechanism is also considered to be present in all subsequent scenarios due to the gene topology in the native GBL system. [2] [3]
- RA – The formation of a complex between ScbA and ScbR proteins (ScbA-ScbR), which relieves ScbR repression, while at the same time activates the transcription of scbA, and in turn of SCBs. [1]
- AS – The antisense effect conferred by convergent transcription of the scbR and scbA genes. In this case, transcripts with a segment of complementary sequence may lead to interactions between sense-antisense full length transcripts of the two genes, thus leading to the formation of a fast degrading complex of the two mRNAs and inhibition of translation. [2]
- Ract – The potential dual role of ScbR protein, which acts as a repressor for its own gene and as an activator for scbA [4]
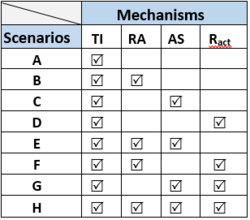
The 8 simulated scenarios (A-H) include different combinations of the four investigated mechanisms (Table on the right).
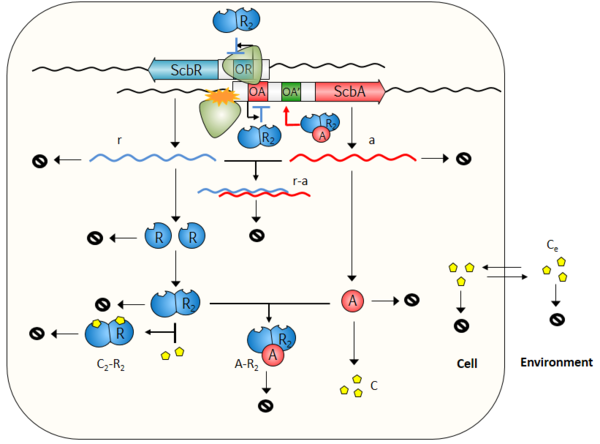
The model consists of two compartments, the Cell and the Environment. A schematic description of the model is provided below. More information can be obtained by clicking on each reaction arrow in the figure.
The model scripts for the different scenarios can be found in pdf form here: File:GBL Models.pdf
Species
| Name | Description | Compartment |
|---|---|---|
| OR | Operator site of ScbR upstream scbR promoter | Cell |
| OA | Operator site of ScbR within scbA promoter as well as putative operator site OA' of AR2 complex | Cell |
| OR-R2 | Complex of R protein and operator site upstream scbR promoter | Cell |
| OR-2R2 | Complex of two R proteins and operator site upstream scbR promoter | Cell |
| OA-R2 | Complex of R protein and operator site within scbA promoter | Cell |
| OA-2R2 | Complex of two R proteins and operator site within scbA promoter | Cell |
| OA-AR2 | Complex of putative operator site presumed to activate scbA gene and AR2 | Cell |
| OA-R2-AR2 | Complex of operator site within scbA promoter with AR2 and R2 | Cell |
| OA-2R2-AR2 | Complex of operator site within scbA promoter with AR2 and two R2 | Cell |
| r | mRNA transcript of scbR gene | Cell |
| a | mRNA transcript of scbA gene | Cell |
| r-a | Complex of full length scbR and scbA mRNAs due to antisense effect | Cell |
| R | ScbR protein | Cell |
| R2 | ScbR homo-dimer | Cell |
| A | ScbA protein | Cell |
| C | SCBs (γ-butyrolactones) | Cell |
| AR2 | ScbA-ScbR complex | Cell |
| S | Glycerol derivative and β-keto acid derivative precursors | Cell |
| C2-R2 | SCBs-ScbR complex | Cell |
| Ce | Extracellular SCBs (γ-butyrolactones) | Environment |
Information on the initial concentrations of all species can be found in the relevant section of the Model Overview.
Reactions
The reactions per compartment are the following:
|
Cell |
Cell-Environment
Environment |
Parameter Overview
An overview for all the parameters of the model can be found in the Model Overview.
In order to analyse the system, uncertainty modelling is employed. This means that each parameter is described by a probability distribution which expresses the associated uncertainty. The methods employed to generate the probability distributions are described in Quantification of parameter uncertainty.
Stochastic aspects
Stochasticity is an inherent trait of gene expression (fluctuations in transcription and translation) and cannot be predicted or eliminated. When the number of molecules of the system is large, stochastic events can be ignored, however in a system involving a small number of molecules (dozens or hundreds), such as a regulatory or signalling system, it can have a significant impact. The noise can originate from different sources, such as thermal fluctuations concerning individual molecules within the cell (intrinsic noise), randomness in the autoinducer diffusion from the cell to the environment, randomness on gene transcription and protein synthesis, or can even be caused by external sources (extrinsic noise) in the cell's environment.
In order to study the role of noise in a population of Streptomyces communicating via 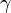 -butyrolactones, a stochastic model developed by Weber et al. [5] which is simulating the bacterial growth in a colony is employed. The model considers each bacterium as an individual cell carrying a copy of the GBL regulatory network. The resulting ensemble of all reactions in all cells is functioning as one global system. The cells are coupled through the autoinducer diffusion reaction as follows: Each time a GBL molecule diffuses from out of one cell into the external environment it increases the number of molecules in the medium by one, thereby also increasing the probability of a GBL molecule to diffuse into another cell of the colony. More information on the stochastic bacterial colony growth simulation can be found in the section Dynamics of bacterial colony growth.
-butyrolactones, a stochastic model developed by Weber et al. [5] which is simulating the bacterial growth in a colony is employed. The model considers each bacterium as an individual cell carrying a copy of the GBL regulatory network. The resulting ensemble of all reactions in all cells is functioning as one global system. The cells are coupled through the autoinducer diffusion reaction as follows: Each time a GBL molecule diffuses from out of one cell into the external environment it increases the number of molecules in the medium by one, thereby also increasing the probability of a GBL molecule to diffuse into another cell of the colony. More information on the stochastic bacterial colony growth simulation can be found in the section Dynamics of bacterial colony growth.
The system of chemical reactions is simulated by employing the Gillespie algorithm [6] which computes the propensities of all species according to the volume of the cell at a given time and, pursuant to them, determines the time of the next reaction, chooses a reaction channel among all possible reactions and updates the number of molecules according to the reaction stoichiometry. It should be noted that even though in principle the Gillespie algorithm should be adapted in order to take into account the time-dependent change in the cell volume, this is not done as the rates of the internal system are faster than the rate of variation of the cell volume. This means that the volume increase is negligible during the time interval until the next reaction and therefore we can assume that the volume-dependent propensities remain constant until the next reaction occurs.
References
- ↑ 1.0 1.1 S. Mehra, S. Charaniya, E. Takano, and W.-S. Hu. A bistable gene switch for antibiotic biosynthesis: The butyrolactone regulon in streptomyces coelicolor. PLoS ONE, 3(7), 2008.
- ↑ 2.0 2.1 2.2 A. Chatterjee, L. Drews, S. Mehra, E. Takano, Y.N. Kaznessis, and W.-S. Hu. Convergent transcription in the butyrolactone regulon in streptomyces coelicolor confers a bistable genetic switch for antibiotic biosynthesis. PLoS ONE, 6(7), 2011.
- ↑ E. Takano. γ-butyrolactones: Streptomyces signalling molecules regulating antibiotic production and differentiation. Current Opinion in Microbiology, 9(3):287–294, 2006.
- ↑ E. Takano, R. Chakraburtty, T. Nihira, Y. Yamada, & M. J. Bibb. A complex role for the γ‐butyrolactone SCB1 in regulating antibiotic production in Streptomyces coelicolor A3 (2). Molecular microbiology 41(5): 1015-1028,2001.
- ↑ M. Weber and J. Buceta. Dynamics of the quorum sensing switch: Stochastic and non-stationary effects. BMC Systems Biology 2013,(7):6
- ↑ Gillespie D.T. Exact stochastic simulation of coupled chemical reactions. Journal of Physical Chemistry 1977,81(25):2340–2361